Wolfram Function Repository
Instant-use add-on functions for the Wolfram Language
Function Repository Resource:
Compute the rigid-body geometric parameters for a pair of nucleic acid residues in a biomolecule
ResourceFunction["NucleicAcidBasePairParameters"][biomol,{{model,chainID1,residueID1},{model,chainID2,residueID2}}] gives an association of the base-pair parameters for residueID1 in chainID1 with residueID2 in chainID2 of model of the DNA or RNA BioMolecule biomol. | |
ResourceFunction["NucleicAcidBasePairParameters"][biomol,{{chainID1,residueID1},{chainID2,residueID2}}] is equivalent to ResourceFunction["NucleicAcidBasePairParameters"][biomol,{{1,chainID1,residueID1},{1,chainID2,residueID2}}]. | |
ResourceFunction["NucleicAcidBasePairParameters"][biomol,{{{m1,c11,r11},{m1,c12,r12}},{{m2,c21,r21},{m2,c22,r22}},…}] gives an association of base-pair parameter associations. |
| "BasePair" | nucleotides and relative orientation of the base planes (+/-) |
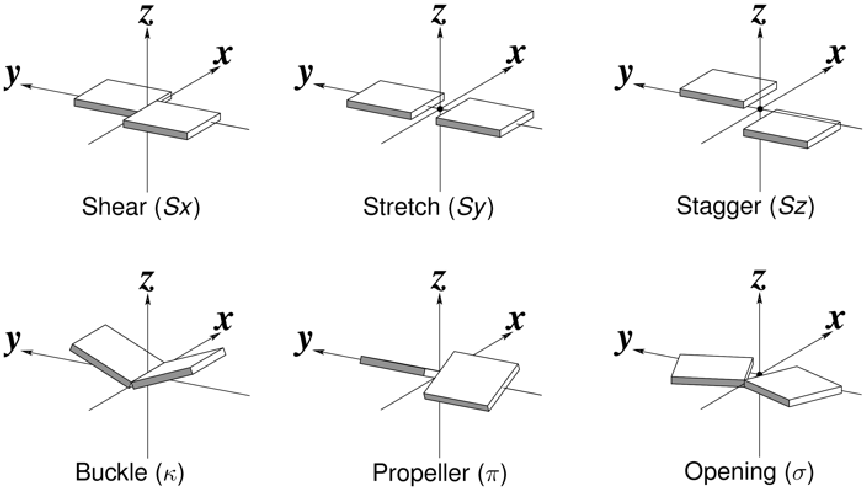
| "Shear" | opposite displacement of the bases in the x direction |
| "Stretch" | opposite displacement of the bases in the y direction |
| "Stagger" | opposite displacement of the bases in the z direction |
| "Buckle" | disrotatory rotation of the bases about the x axis |
| "Propeller" | disrotatory rotation of the bases about the y axis |
| "Opening" | disrotatory rotation of the bases about the z axis |
| "Angle" | angle between the normals of the base planes |
Import the structure for the yeast phenylalanine tRNA:
| In[1]:= |
| Out[1]= |
Show the bases U6 and A67 that have a conventional Watson-Crick geometry:
| In[2]:= |
| Out[2]= | 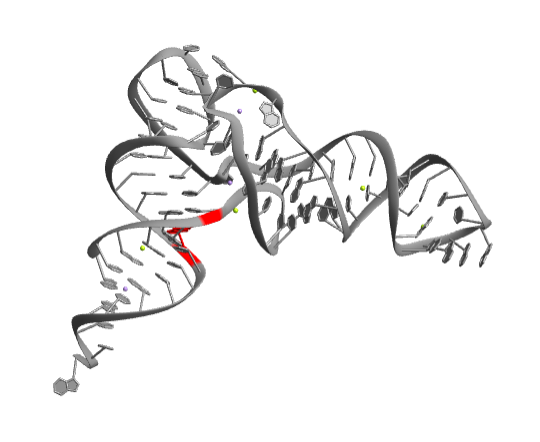 |
Compute the base-pair parameters for residues U6 and A67:
| In[3]:= |
| Out[3]= | 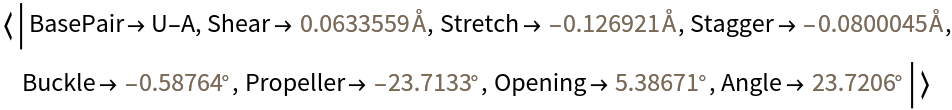 |
Compute the parameters for several base-pairs:
| In[4]:= |
| Out[4]= |  |
Import a dodecamer B-DNA structure:
| In[5]:= |
| Out[5]= |
Highlight the bases A-DC1 and B-DG12:
| In[6]:= |
| Out[7]= | 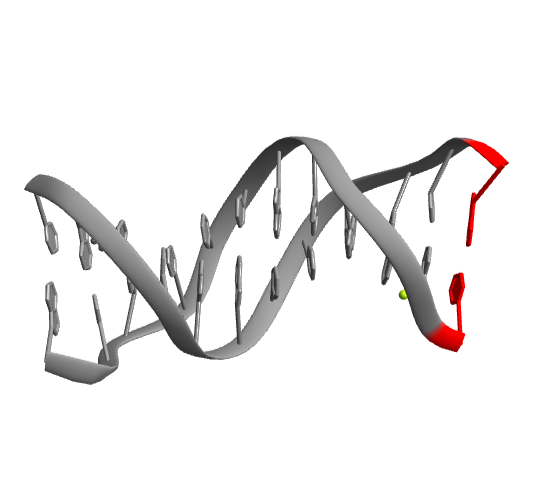 |
Compute the parameters for residues A-DC1 and B-DG12:
| In[8]:= |
| Out[8]= | 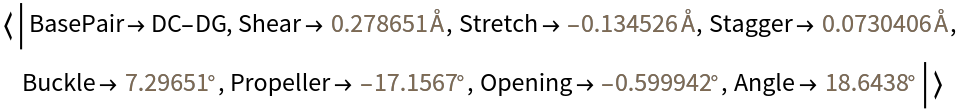 |
Import a short RNA that has four models:
| In[9]:= |
Highlight the residues A-G1 and A-C16 in each:
| In[10]:= | ![BioMoleculePlot3D[#, ImageSize -> Small, ColorRules -> {{"A", 1 | 16} -> StandardRed, _ -> StandardGray}, ImageSize -> Small] & /@ biomol["Models"] // Partition[#, UpTo[2]] & // Grid](https://www.wolframcloud.com/obj/resourcesystem/images/58b/58ba23eb-2c5b-46cb-b6ec-2ddd2246fe70/78f052c0d3ff9a3b.png) |
| Out[11]= |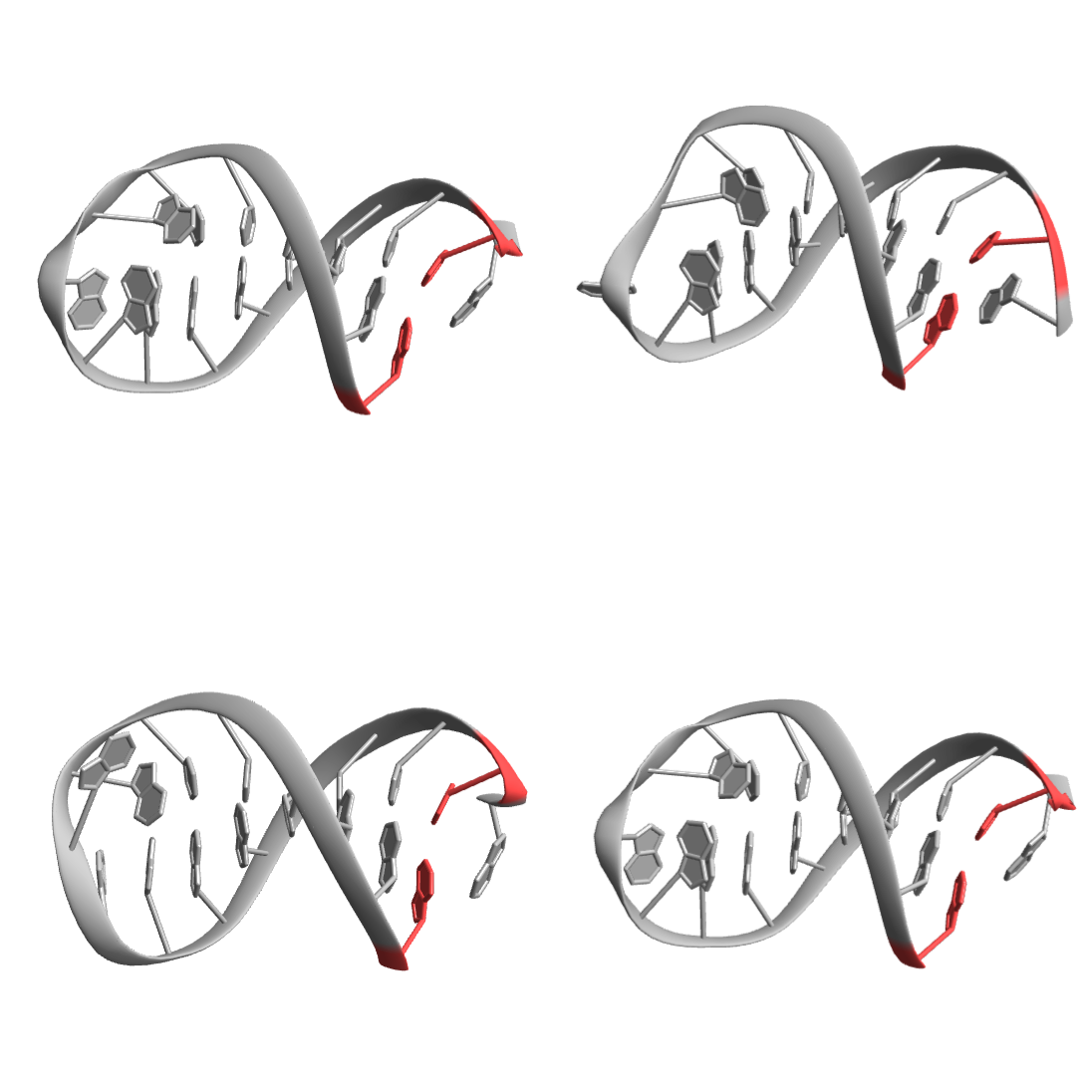  |
Compute the parameters for the residues A-G1 and A-C16 for all the models:
| In[12]:= |
| Out[12]= |  |
Base-pairing in duplex DNA is characteristically different from that in duplex RNA.
Import a duplex DNA:
| In[13]:= | ![(duplexDNA = BioMolecule[ExternalIdentifier["PDBStructureID", "355D"]]) // BioMoleculePlot3D[#, ImageSize -> Small] &](https://www.wolframcloud.com/obj/resourcesystem/images/58b/58ba23eb-2c5b-46cb-b6ec-2ddd2246fe70/144272364094f648.png) |
| Out[13]= |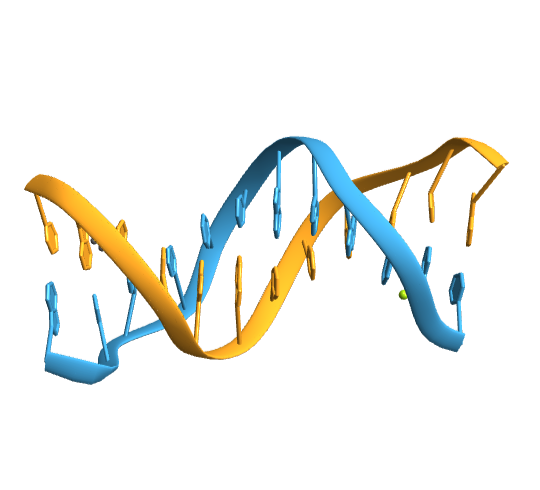  |
Compute the parameters for all the base-pairs:
| In[14]:= | ![residuePairs = Thread@MapThread[
Thread[{##}] &, {{"A", "B"}, MapAt[Reverse, Values@duplexDNA["ResidueIDs"][[{"A", "B"}]], 2]}];
basePairParameters = ResourceFunction["NucleicAcidBasePairParameters"][duplexDNA, residuePairs];
bpDataDNA = KeyValueMap[Prepend[#2, "Residues" -> #1] &, basePairParameters] // Tabular](https://www.wolframcloud.com/obj/resourcesystem/images/58b/58ba23eb-2c5b-46cb-b6ec-2ddd2246fe70/50adc046dbbf2c84.png) |
| Out[15]= | 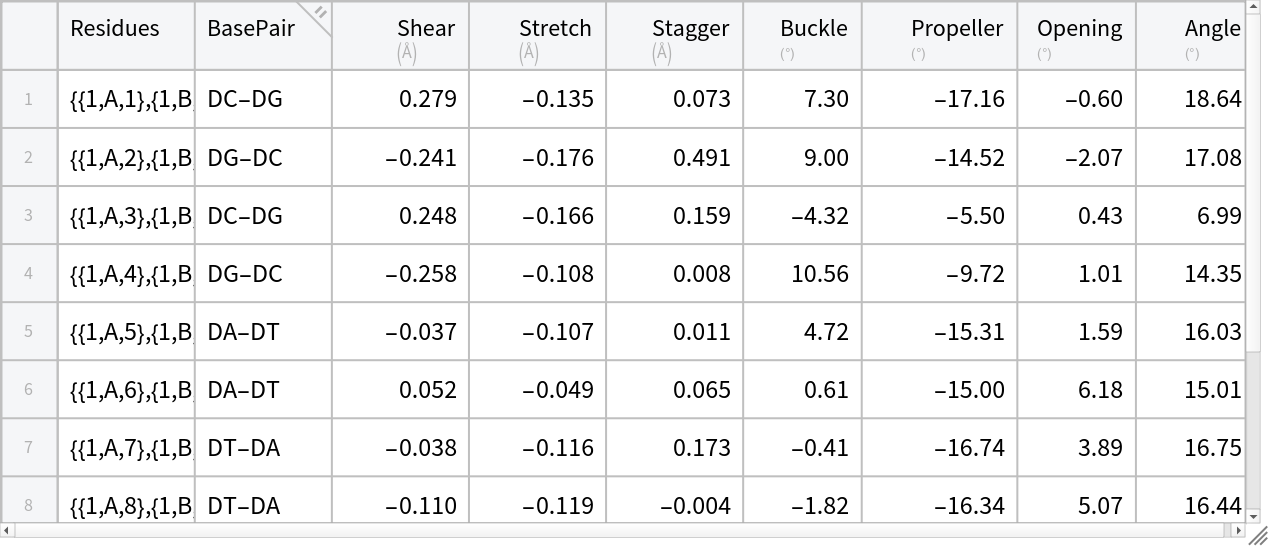 |
Import a duplex RNA:
| In[16]:= | ![(duplexRNA = BioMolecule[ExternalIdentifier["PDBStructureID", "4JRD"]]) // BioMoleculePlot3D[#, ImageSize -> Small] &](https://www.wolframcloud.com/obj/resourcesystem/images/58b/58ba23eb-2c5b-46cb-b6ec-2ddd2246fe70/27b30bd32c105d14.png) |
| Out[16]= | 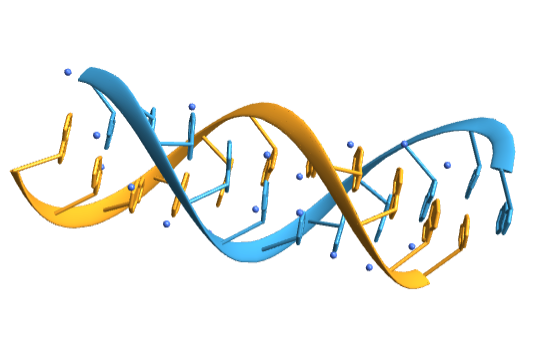 |
Compute the parameters for all the base-pairs:
| In[17]:= | ![residuePairs = Thread@MapThread[
Thread[{##}] &, {{"A", "B"}, MapThread[
Construct, {{Rest, Most}, Values@duplexRNA["ResidueIDs"][[{"A", "B"}]]}]}];
basePairParameters = ResourceFunction["NucleicAcidBasePairParameters"][duplexRNA, residuePairs];
bpDataRNA = KeyValueMap[Prepend[#2, "Residues" -> #1] &, basePairParameters] // Tabular](https://www.wolframcloud.com/obj/resourcesystem/images/58b/58ba23eb-2c5b-46cb-b6ec-2ddd2246fe70/6610889e5314398c.png) |
| Out[18]= | 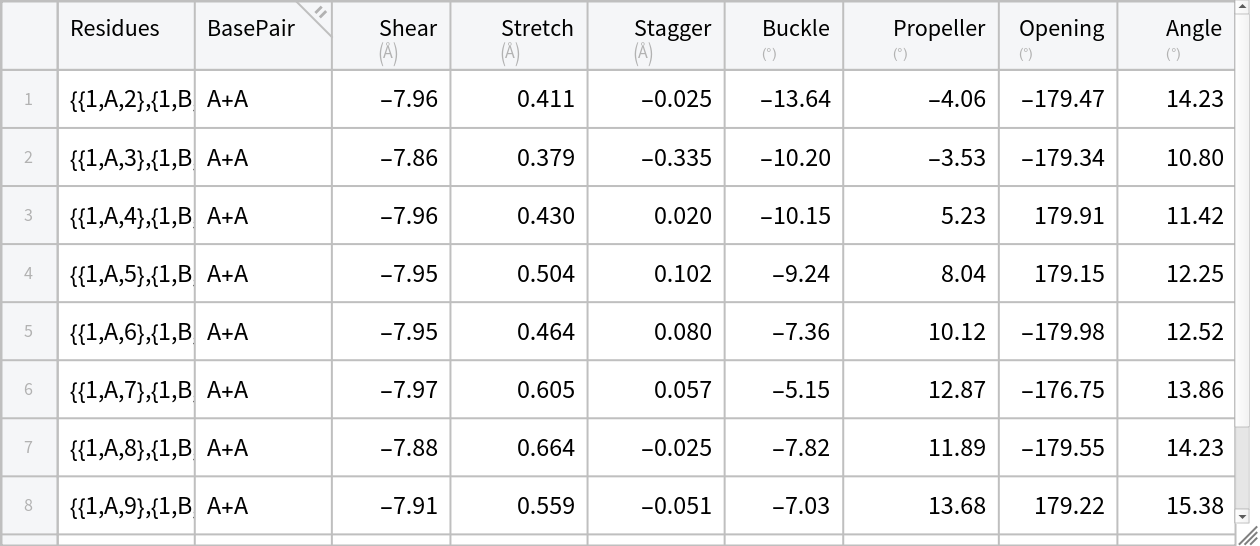 |
Plot the shear and stretch parameters for the two helices:
| In[19]:= | ![ListPlot[
Normal /@ {bpDataDNA[All, {"Shear", "Stretch"}], bpDataRNA[All, {"Shear", "Stretch"}]}, PlotRange -> {{-10, 2}, {-2, 2}}, AspectRatio -> Automatic, Axes -> True, Frame -> True, FrameLabel -> {"Shear", "Stretch"}, PlotLegends -> {"B-BNA", "Poly-A RNA"}]](https://www.wolframcloud.com/obj/resourcesystem/images/58b/58ba23eb-2c5b-46cb-b6ec-2ddd2246fe70/2acc4e7e626d6023.png) |
| Out[19]= | 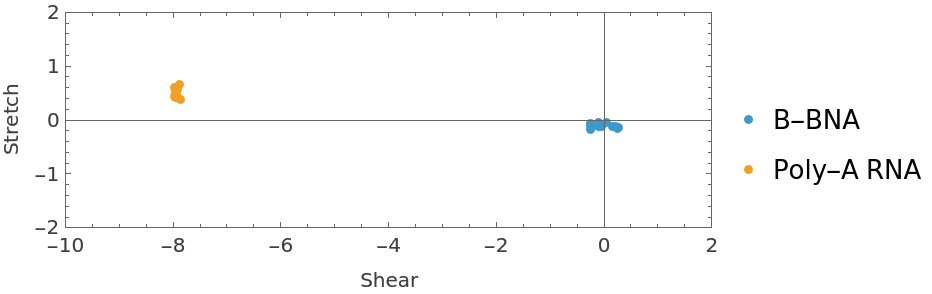 |
Plot the propeller and buckle parameters for the two helices:
| In[20]:= | ![ListPlot[
Normal /@ {bpDataDNA[All, {"Propeller", "Buckle"}], bpDataRNA[All, {"Propeller", "Buckle"}]}, PlotRange -> {{-20, 20}, {-20, 20}}, AspectRatio -> Automatic, Axes -> True, Frame -> True, FrameLabel -> {"Propeller", "Buckle"}, PlotLegends -> {"B-BNA", "Poly-A RNA"}]](https://www.wolframcloud.com/obj/resourcesystem/images/58b/58ba23eb-2c5b-46cb-b6ec-2ddd2246fe70/6c2aff001533b879.png) |
| Out[20]= | 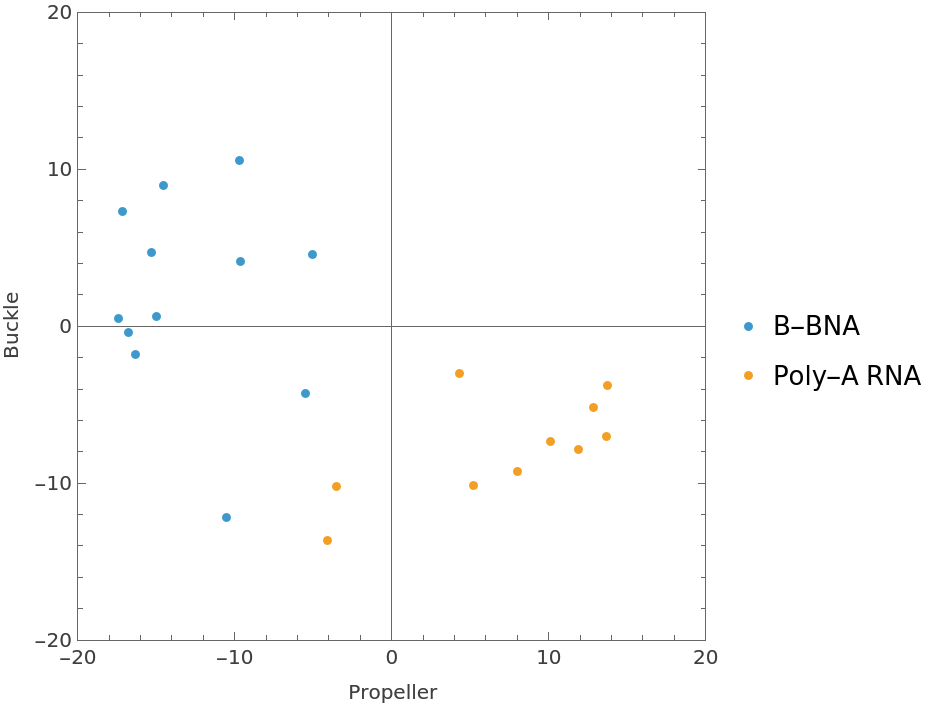 |
This work is licensed under a Creative Commons Attribution 4.0 International License