Wolfram Function Repository
Instant-use add-on functions for the Wolfram Language
Function Repository Resource:
Simulate an aggregation system (a 2D array of cells in which new cells are added at positions with certain neighborhood configurations) as a multiway system
ResourceFunction["MultiwayAggregationSystem"][rule,init,n] generates the results of n steps in the evolution of the multiway aggregation system with the specified rule, starting from initial conditions init. | |
ResourceFunction["MultiwayAggregationSystem"][rule,init,n,"prop"] gives the property "prop" for the specified multiway aggregation system evolution. | |
ResourceFunction["MultiwayAggregationSystem"][rule→sel,init,n,…] uses the function sel to select which of the events obtained at each step to include in the evolution. |
| {n,{2,1},{1,1}} | 9-neighbor totalistic rule |
| {n,{2,{{0,1,0},{1,1,1},{0,1,0}}},{1,1}} | 5-neighbor totalistic rule |
| {n,{2,{{0,2,0},{2,1,2},{0,2,0}}},{1,1}} | 5-neighbor outer totalistic rule |
| "TotalisticCode" | n | totalistic code |
| "OuterTotalisticCode" | n | outer totalistic code |
| "Dimension" | d | overall dimension (must always be 2) |
| "Neighborhood" | type | neighborhood type |
| "Range" | r | range of rule |
| "Colors" | k | number of colors |
| "GrowthCases" | {g1,g2,…} | make a cell 1 when gi of its neighbors are 1 |
| "GrowthSurvivalCases" | {{g1,…},{s1,…}} | 1 for gi neighbors; unchanged for si |
| "GrowthDecayCases" | {{g1,…},{d1,…}} | 1 for gi neighbors; 0 for di |
| 5 or "VonNeumann" | CrossMatrix[1] |
| 9 or "Moore" | BoxMatrix[1] |
| 2D general rules | 2256 |
| 2D 9-neighbor totalistic rules | 210 |
| 2D 5-neighbor totalistic rules | 26 |
| 2D 5-neighbor outer totalistic rules | 210 |
| 2D outer totalistic rules | 217+1 |
| "Sequential" | applies the first possible replacement (sequential substitution system) |
| "Random" | applies a random replacement |
| {"Random",n} | applies n randomly chosen replacements |
| "AllStatesList" | the list of all states generated at each successive step (default) |
| "StatesCountsList" | the number of distinct states generated at each successive step |
| "AllStatesListUnmerged" | the list of all states without any merging |
| "PredecessorRulesList" | the list of states and their corresponding predecessor states at each successive step |
| "EvolutionGraph" | graph formed by the evolution process, with no merging between different time steps |
| "EvolutionGraphStructure" | evolution graph without labeling |
| "EvolutionGraphFull" | graph formed by the evolution process, including equivalent events |
| "EvolutionGraphFullStructure" | full evolution graph without labeling |
| "EvolutionGraphUnmerged" | graph formed by the evolution process, with no merging of equivalent states |
| "EvolutionGraphUnmergedStructure" | unmerged evolution graph without labeling |
| "EvolutionGraphWeighted" | graph formed by the evolution process, with edges weighted by event multiplicity |
| "EvolutionGraphWeightedStructure" | weighted evolution graph without labeling |
| "StatesGraph" | graph of how each distinct state leads to other states |
| "StatesGraphStructure" | states graph without labeling |
| "AllEventsList" | the list of all events that occur at each successive step |
| "EvolutionEventsGraph" | graph showing the evolution process with updating events explicitly included |
| "EvolutionEventsGraphStructure" | evolution events graph without labeling |
| "CausalGraph" | graph of all causal relations between updating events |
| "CausalGraphStructure" | causal graph without labeling |
| "EvolutionCausalGraph" | combined graph of evolution process and causal relationships between events |
| "EvolutionCausalGraphStructure" | evolution causal graph without labeling |
| "CausalGraphInstances" | list of distinct causal graphs for all possible choices of event sequences |
| "CausalGraphStructureInstances" | causal graph instances without labeling |
| "EvolutionCausalGraphInstances" | list of distinct evolution causal graphs for all possible choices of event sequences |
| "EvolutionCausalGraphStructureInstances" | evolution causal graph instances without labeling |
| "BranchPairsList" | list of all branch pairs (i.e. critical pairs) generated in the states graph |
| "NewBranchPairsList" | list of all new branch pairs generated at each successive step |
| "EvolutionBranchPairsList" | list of all branch pairs generated in the evolution graph |
| "NewEvolutionBranchPairsList" | list of all new evolution branch pairs generated at each successive step |
| "BranchPairEventsList" | list of all events yielding branch pairs |
| "NewBranchPairEventsList" | list of all events yielding new branch pairs at each successive step |
| "EvolutionBranchPairEventsList" | list of all events yielding evolution branch pairs |
| "NewEvolutionBranchPairEventsList" | list of all events yielding new evolution branch pairs at each successive step |
| "BranchialGraph" | graph of branch pair ancestry at a given step |
| "BranchialGraphStructure" | branchial graph without labeling |
| "AllStatesBranchialGraph" | graph of branch pair ancestry across all steps |
| "AllStatesBranchialGraphStructure" | all states branchial graph without labeling |
| "EvolutionBranchialGraph" | graph of evolution branch pair ancestry at a given step |
| "EvolutionBranchialGraphStructure" | evolution branchial graph without labeling |
| "AllStatesEvolutionBranchialGraph" | graph of evolution branch pair ancestry across all steps |
| "AllStatesEvolutionBranchialGraphStructure" | all states evolution branchial graph without labeling |
| "EventBranchialGraph" | graph of branch pair event ancestry at a given step |
| "EventBranchialGraphStructure" | event branchial graph without labeling |
| "AllEventsBranchialGraph" | graph of branch pair event ancestry across all steps |
| "AllEventsBranchialGraphStructure" | all events branchial graph without labeling |
| "EvolutionEventBranchialGraph" | graph of evolution branch pair event ancestry at a given step |
| "EvolutionEventBranchialGraphStructure" | evolution event branchial graph without labeling |
| "AllEventsEvolutionBranchialGraph" | graph of evolution branch pair event ancestry across all steps |
| "AllEventsEvolutionBranchialGraphStructure" | all events evolution branchial graph without labeling |
| "BranchPairResolutionsList" | association of all resolved and unresolved branch pairs up to a given step |
| "EvolutionBranchPairResolutionsList" | association of all resolved and unresolved evolution branch pairs up to a given step |
| "CausalInvariantQ" | whether the system is causal invariant (all branch pairs converge) |
| "EvolutionCausalInvariantQ" | whether the system is evolution causal invariant (all evolution branch pairs converge) |
| "KnuthBendixCompletion" | list of Knuth–Bendix completion rules required to force causal invariance |
| "EvolutionKnuthBendixCompletion" | list of Knuth–Bendix completion rules required to force evolution causal invariance |
| "StateWeights" | list of weights for all vertices in the states graph |
| "IncludeStepNumber" | False | whether to label states and events with their respective step numbers |
| "IncludeStateID" | False | whether to label states and events with unique IDs |
| "IncludeInitializationEvents" | False | whether to include pseudoevents that set up initial conditions |
| "IncludeEventInstances" | False | whether to show distinct updating events that connect the same states as separate edges |
| "IncludeStateWeights" | False | whether to weight state vertices by their rate of occurrence at a particular time step |
| "IncludeStatePathWeights" | False | whether to weight state vertices by the number of distinct evolution paths that lead to them |
| "StateRenderingFunction" | Automatic | how to label states that appear in graphs |
| "EventRenderingFunction" | Automatic | how to label events that appear in graphs |
| MaxItems | Infinity | how many instances of a causal graph or evolution causal graph to return |
| "GivePredecessors" | False | whether to label branch pairs with their predecessor state |
| "GiveResolvents" | False | whether to label branch pairs with their resolvent state |
| "IncludeSelfPairs" | False | whether to include trivial branch pairs |
| "IncludeFullBranchialSpace" | False | whether to show all possible states in a given branchial graph |
| "LineThickness" | 1 | absolute line thickness for graph edges |
| Inherited | use the explicit vertex name as the label |
| None | use no label for the vertex |
| "string" | use a shape from the VertexShapeFunction collection |
| func | apply the function func to the name of the vertex |
Show basic multiway aggregation system evolution (for outer totalistic code 4):
| In[1]:= |
| Out[1]= |  |
For totalistic code 18:
| In[2]:= |
| Out[2]= |  |
Generate a graph showing how each state is obtained from the others:
| In[3]:= |
| Out[3]= | 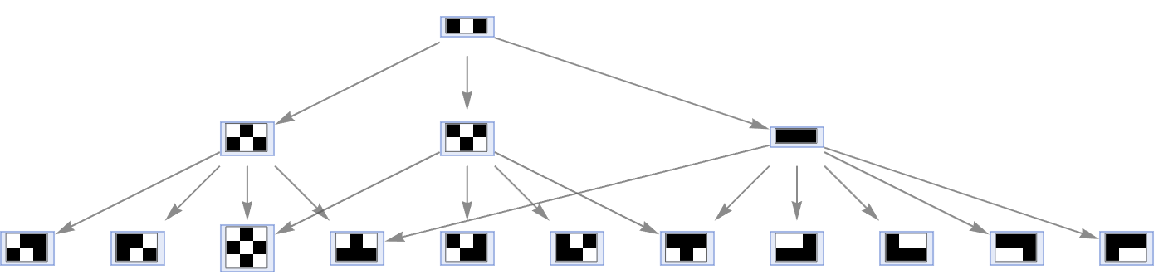 |
Show the structure of the graph, without labels:
| In[4]:= |
| Out[4]= | 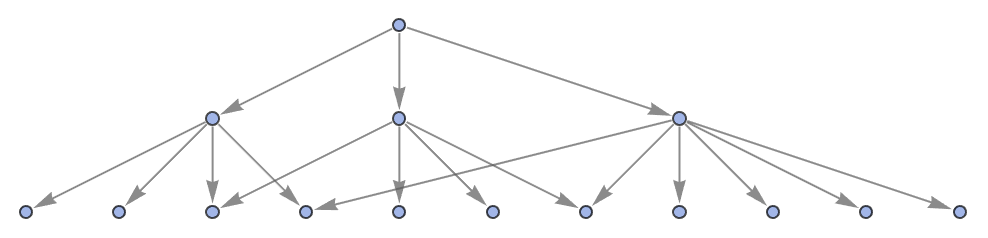 |
Generate the list of all updating events applied at each step:
| In[5]:= |
| Out[5]= | 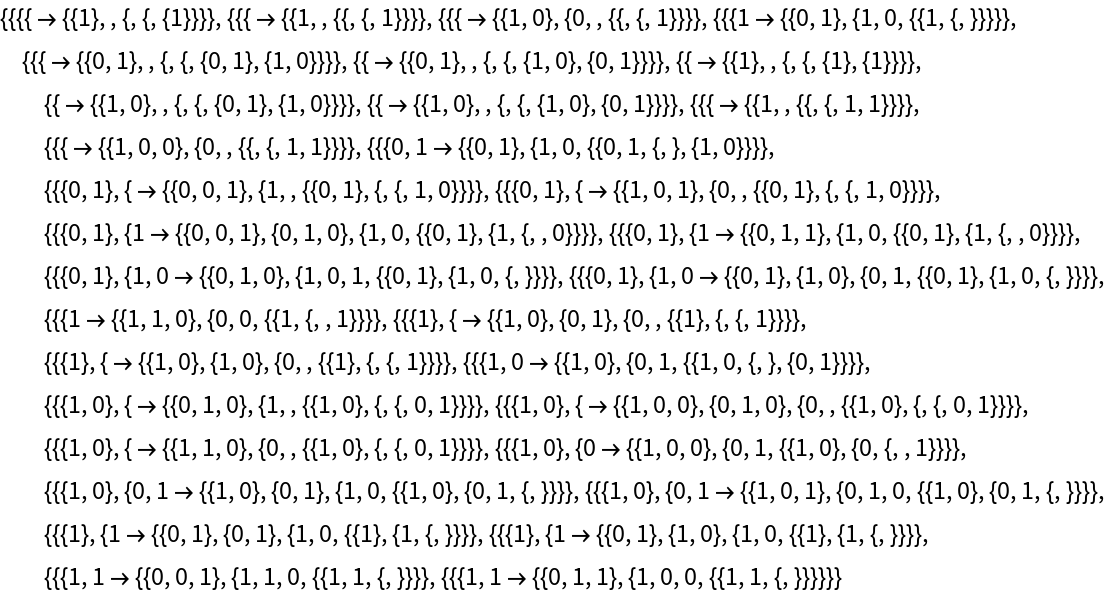 |
Generate a graph of the evolution history, with updating events included:
| In[6]:= |
| Out[6]= | 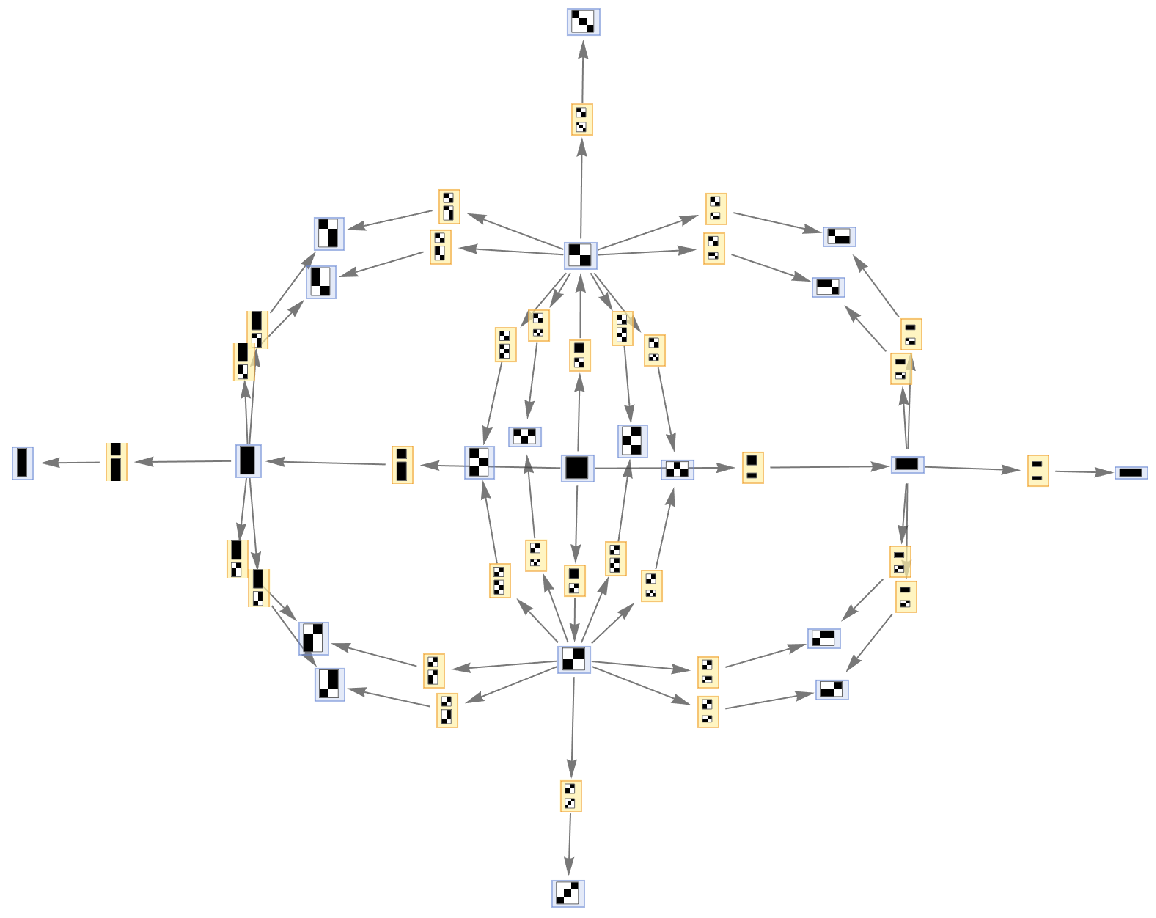 |
Show the structure of the graph, without labels:
| In[7]:= |
| Out[7]= | 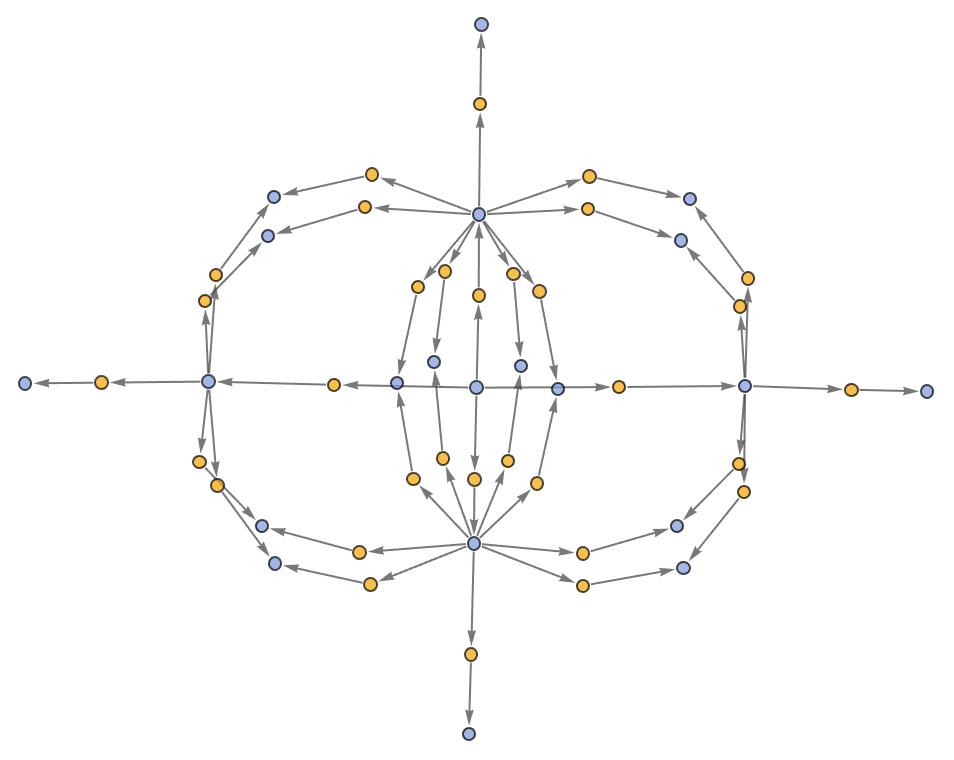 |
Generate the causal graph, showing dependencies between updating events:
| In[8]:= |
| Out[8]= | 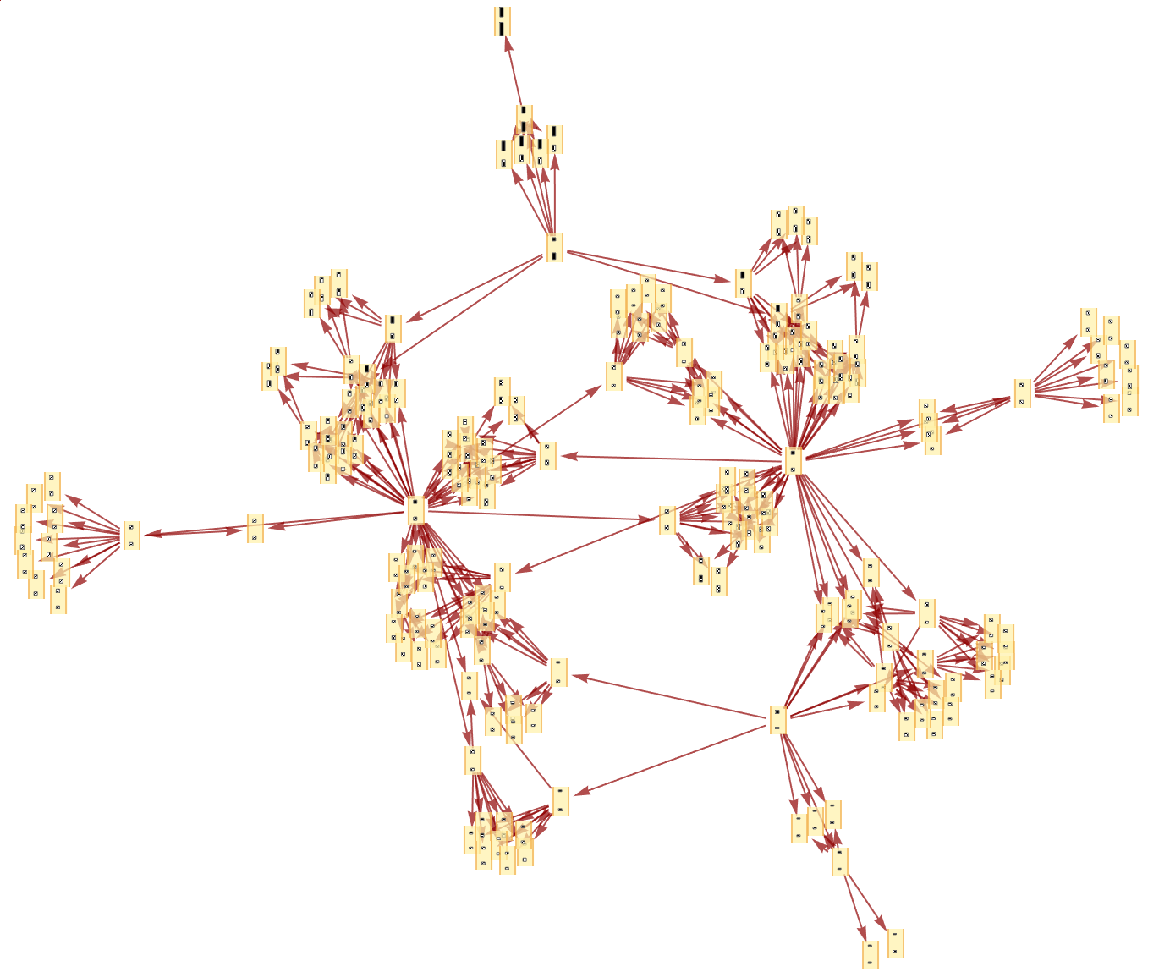 |
Show the structure of the graph, without labels:
| In[9]:= |
| Out[9]= | 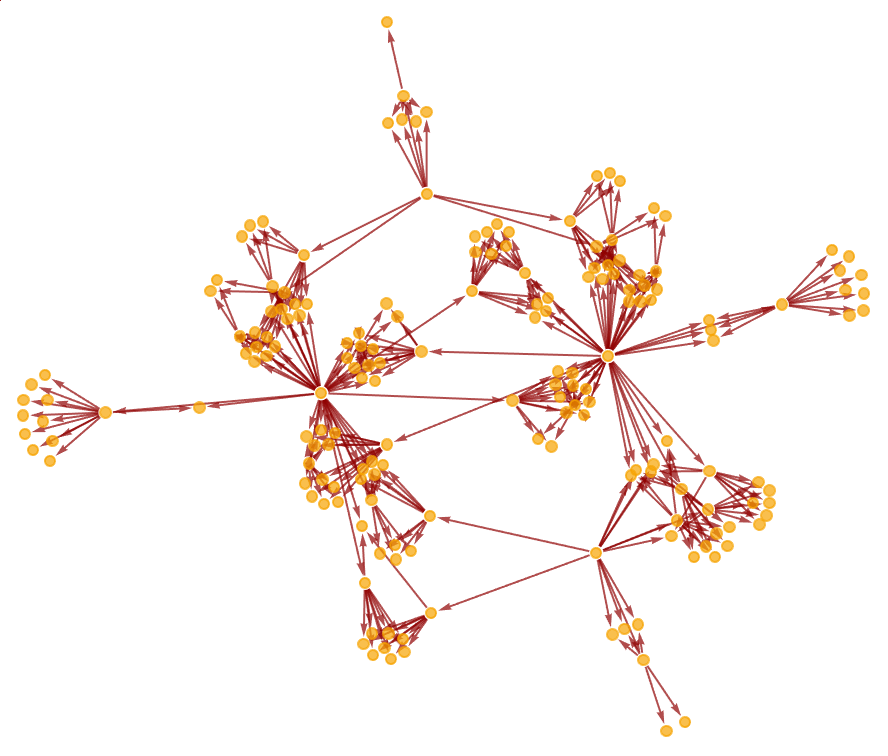 |
Generate the evolution events graph, with causal connections included:
| In[10]:= |
| Out[10]= | 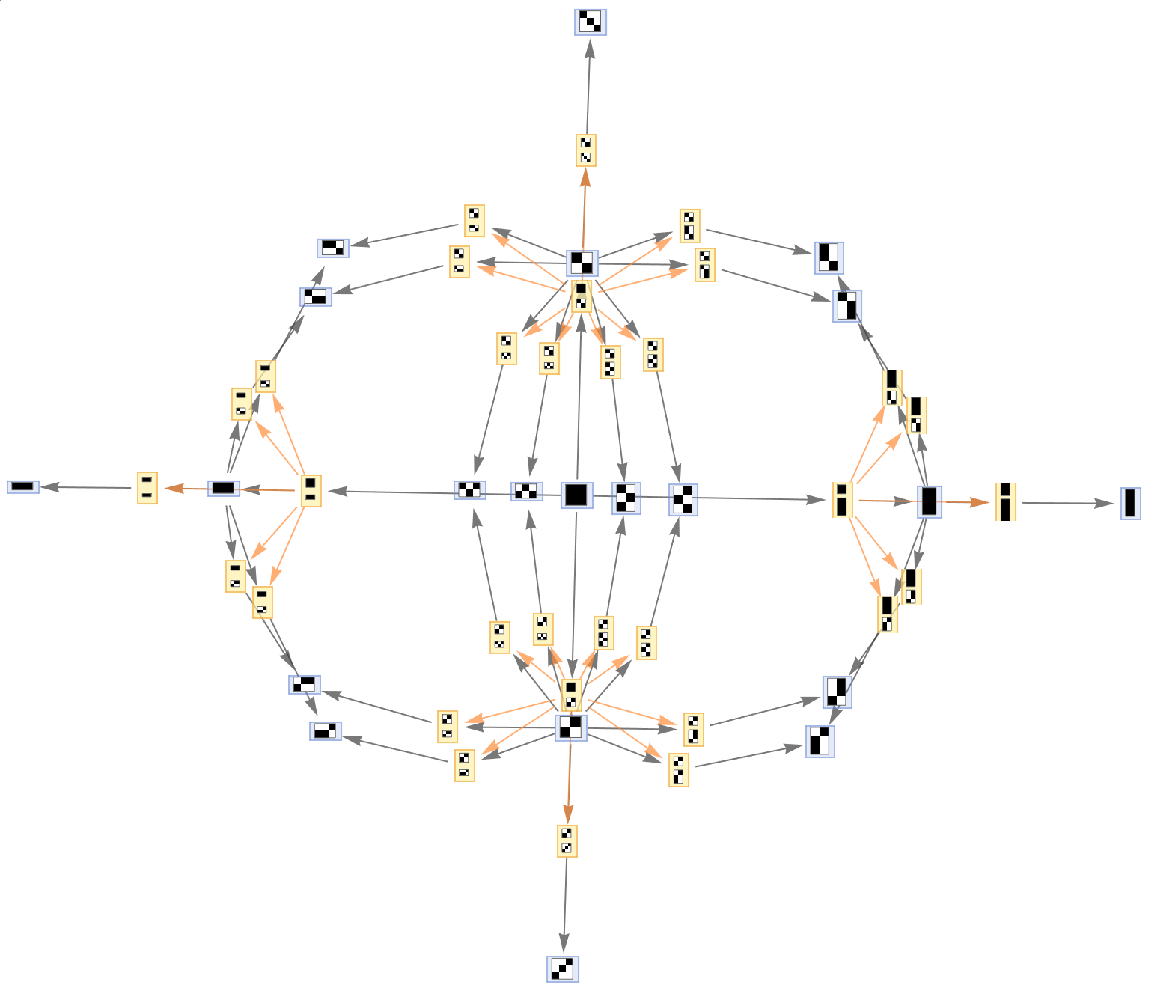 |
Show the structure of the graph, without labels:
| In[11]:= |
| Out[11]= | 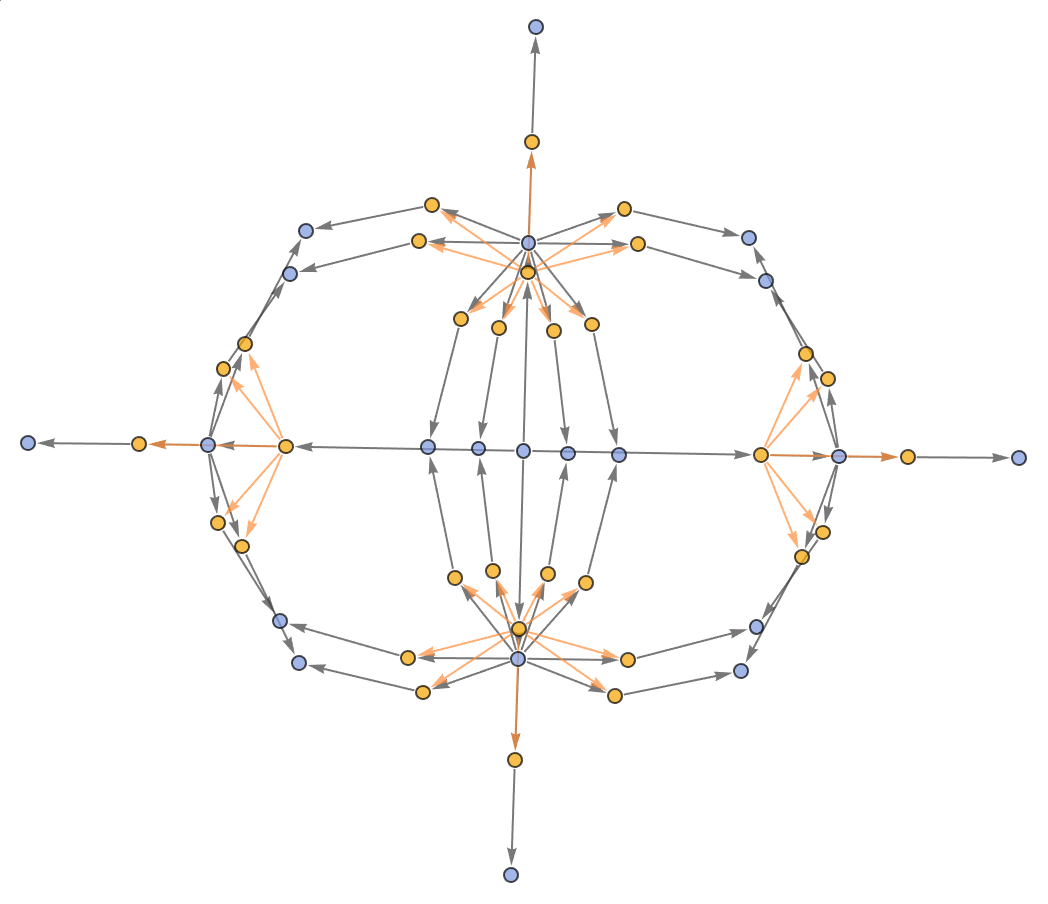 |
Specify an event selection function that picks a random event at each step:
| In[12]:= |
| Out[12]= |  |
Generate causal graphs for all possible choices of event sequences:
| In[13]:= | ![Graph[#, VertexSize -> 1] & /@ Take[ResourceFunction[
"MultiwayAggregationSystem"][<|"OuterTotalisticCode" -> 4, "Dimension" -> 2|>, CenterArray[1, {1, 1}], 3, "CausalGraphInstances"], 10]](https://www.wolframcloud.com/obj/resourcesystem/images/fa5/fa59000f-249d-45f0-bd3b-38f71609dbde/1c9f6970a305e39e.png) |
| Out[13]= | 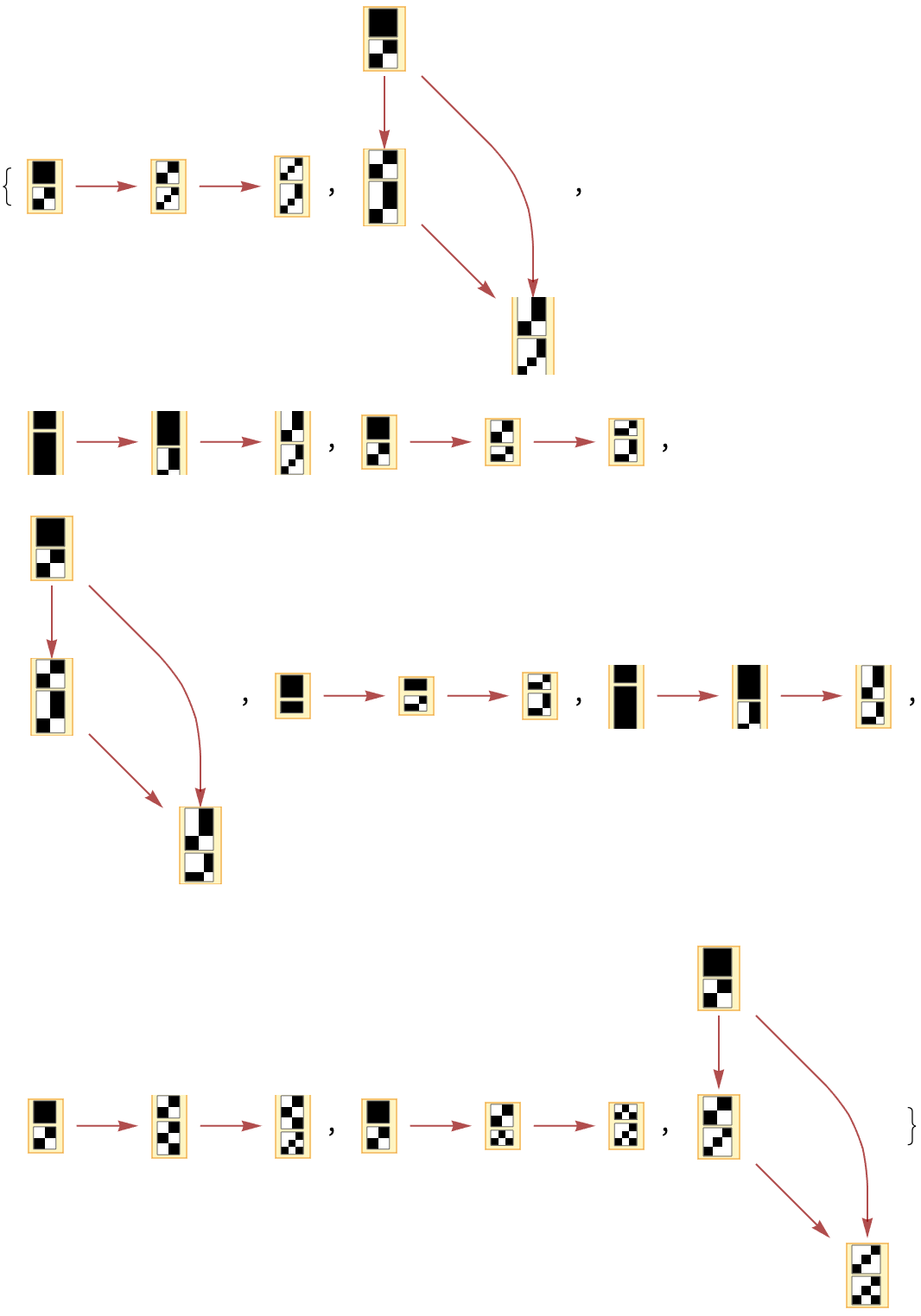 |
Show the structures of the graphs, without labels:
| In[14]:= | ![Take[ResourceFunction[
"MultiwayAggregationSystem"][<|"OuterTotalisticCode" -> 4, "Dimension" -> 2|>, CenterArray[1, {1, 1}], 3, "CausalGraphStructureInstances"], 10]](https://www.wolframcloud.com/obj/resourcesystem/images/fa5/fa59000f-249d-45f0-bd3b-38f71609dbde/6ef15cb6c18b3377.png) |
| Out[14]= | 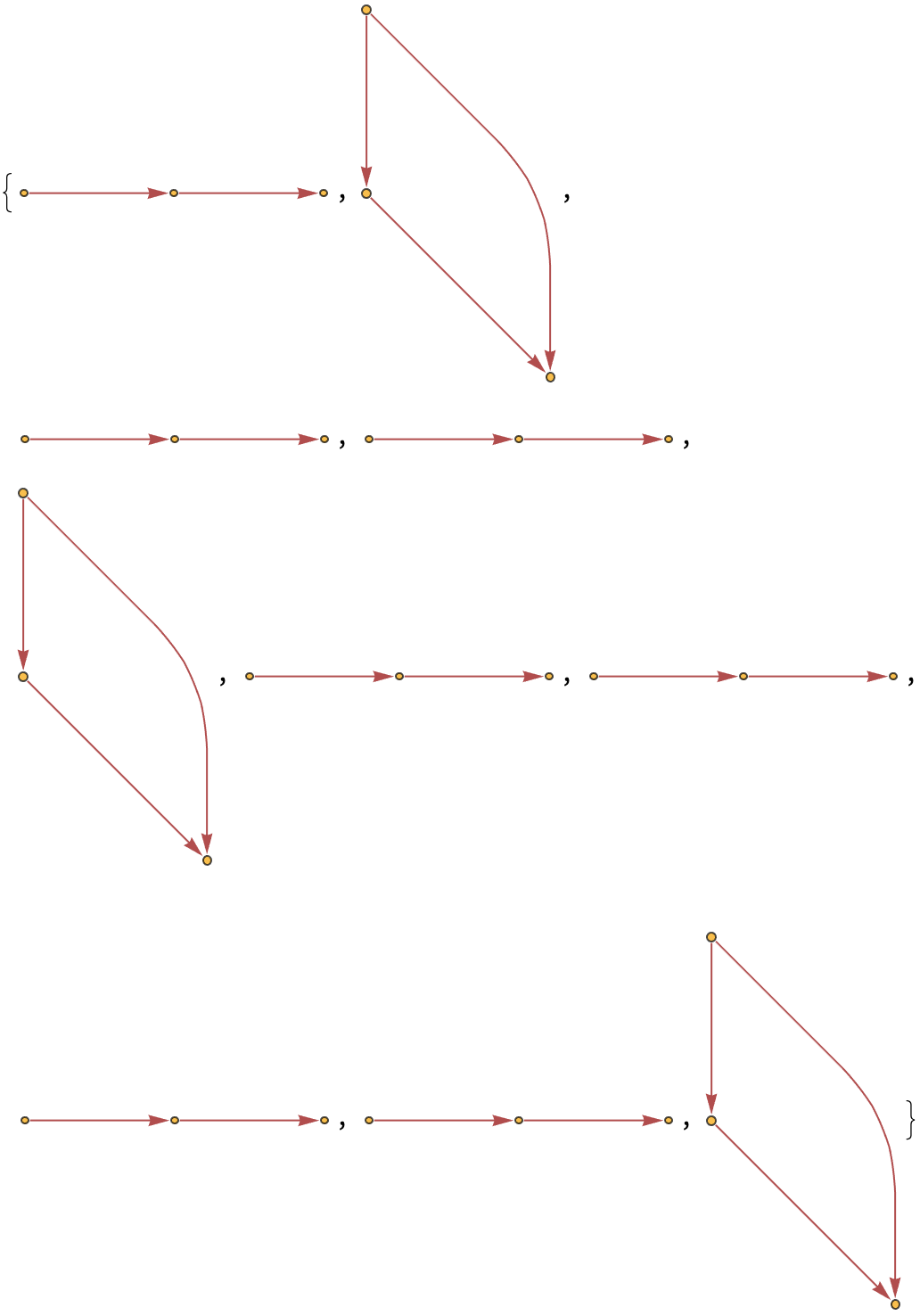 |
Generate the list of all branch pairs (i.e. critical pairs):
| In[15]:= |
| Out[15]= |  |
Generate the association showing which branch pairs converged and which did not:
| In[16]:= |
| Out[16]= |  |
Prove that the system is causal invariant:
| In[17]:= |
| Out[17]= |
Generate a graph showing branch pair ancestry:
| In[18]:= |
| Out[18]= | 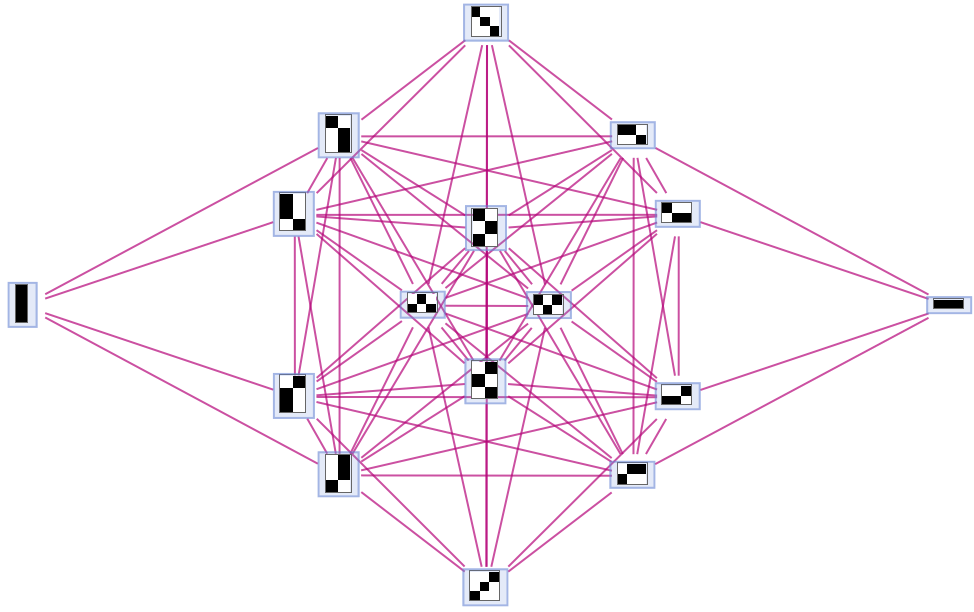 |
Show the structure of the graph, without labels:
| In[19]:= |
| Out[19]= | 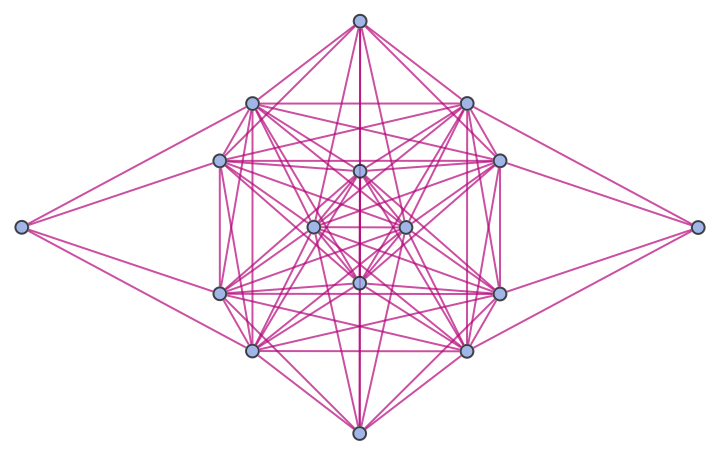 |
Generate a graph showing branch pair event ancestry:
| In[20]:= |
| Out[20]= | 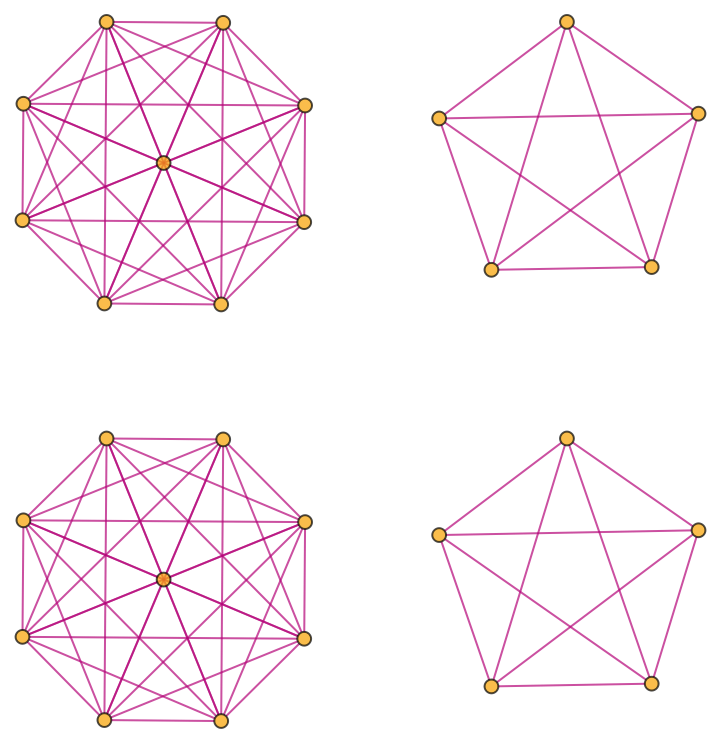 |
Prevent identical states from being merged by including step numbers and state IDs:
| In[21]:= | ![ResourceFunction[
"MultiwayAggregationSystem"][<|"OuterTotalisticCode" -> 466, "Dimension" -> 2|>, {{{1, 0, 1}}}, 2, "StatesGraph", "IncludeStepNumber" -> True, "IncludeStateID" -> True, VertexSize -> 1]](https://www.wolframcloud.com/obj/resourcesystem/images/fa5/fa59000f-249d-45f0-bd3b-38f71609dbde/126be7524bdcacf2.png) |
| Out[21]= | 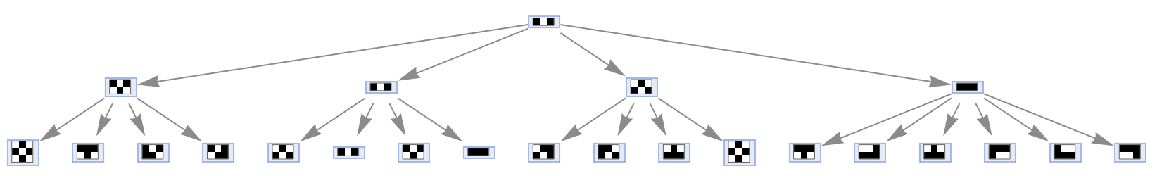 |
| In[22]:= |
| Out[22]= |  |
Generate a graph of full evolution history, with all events included:
| In[23]:= |
| Out[23]= | 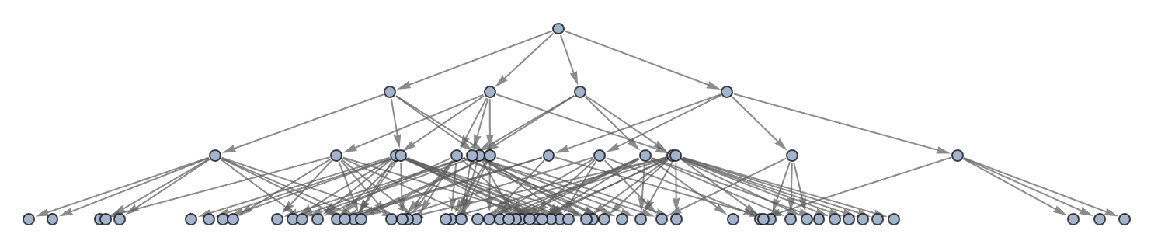 |
Generate a graph of full evolution history, with no merging of equivalent states:
| In[24]:= |
| Out[24]= | 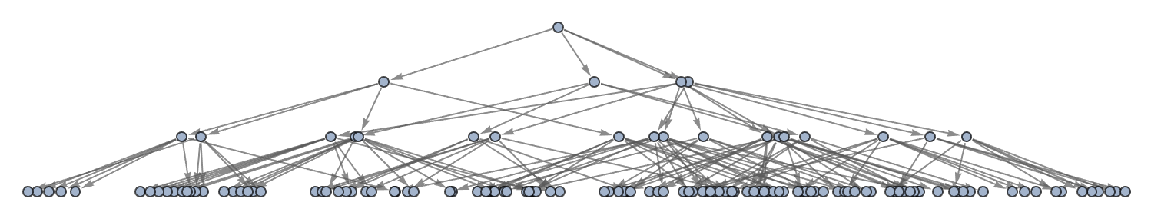 |
Generate a graph of evolution history, with edges weighted by event multiplicity:
| In[25]:= |
| Out[25]= | 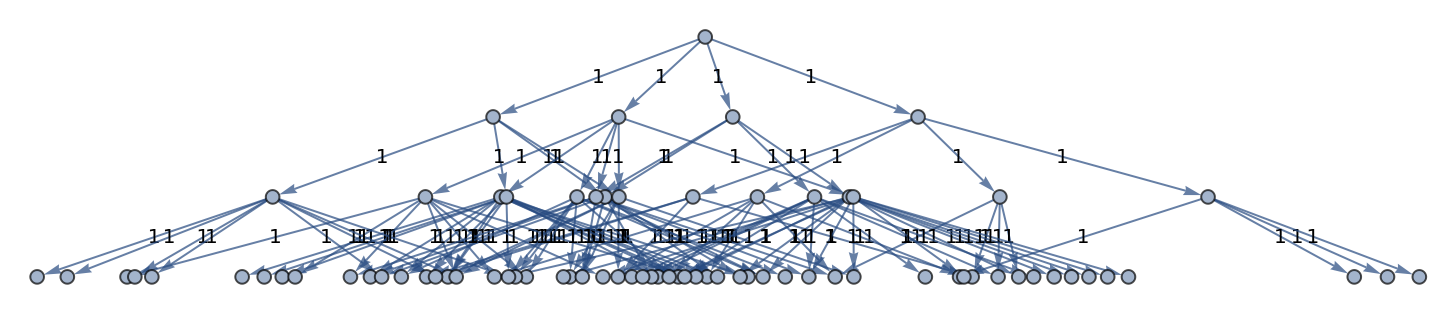 |
Generate a states graph with vertices weighted by their rate of occurrence on each time step:
| In[26]:= | ![ResourceFunction[
"MultiwayAggregationSystem"][<|"OuterTotalisticCode" -> 4, "Dimension" -> 2|>, CenterArray[1, {1, 1}], 2, "StatesGraph", "IncludeStateWeights" -> True, VertexSize -> 1, VertexLabels -> "VertexWeight"]](https://www.wolframcloud.com/obj/resourcesystem/images/fa5/fa59000f-249d-45f0-bd3b-38f71609dbde/4ac01dde3ca2f35d.png) |
| Out[26]= | 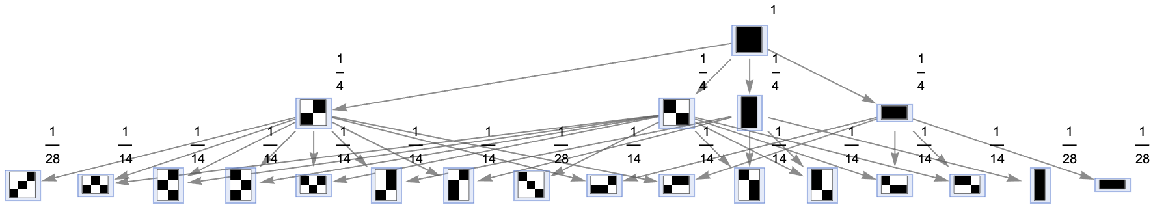 |
Show the structure of the graph, without labels:
| In[27]:= | ![ResourceFunction[
"MultiwayAggregationSystem"][<|"OuterTotalisticCode" -> 4, "Dimension" -> 2|>, CenterArray[1, {1, 1}], 2, "StatesGraphStructure", "IncludeStateWeights" -> True, VertexLabels -> "VertexWeight"]](https://www.wolframcloud.com/obj/resourcesystem/images/fa5/fa59000f-249d-45f0-bd3b-38f71609dbde/4dde89606b485189.png) |
| Out[27]= | 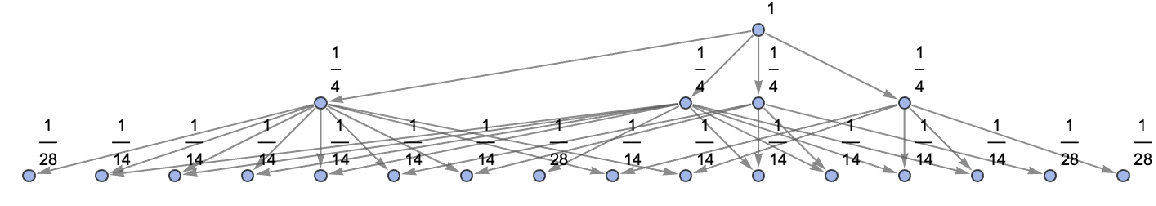 |
Generate a states graph with vertices weighted by the number of distinct evolution paths that lead to them:
| In[28]:= | ![ResourceFunction[
"MultiwayAggregationSystem"][<|"OuterTotalisticCode" -> 4, "Dimension" -> 2|>, CenterArray[1, {1, 1}], 2, "StatesGraph", "IncludeStatePathWeights" -> True, VertexSize -> 1, VertexLabels -> "VertexWeight"]](https://www.wolframcloud.com/obj/resourcesystem/images/fa5/fa59000f-249d-45f0-bd3b-38f71609dbde/70ce0607ba7723be.png) |
| Out[28]= | 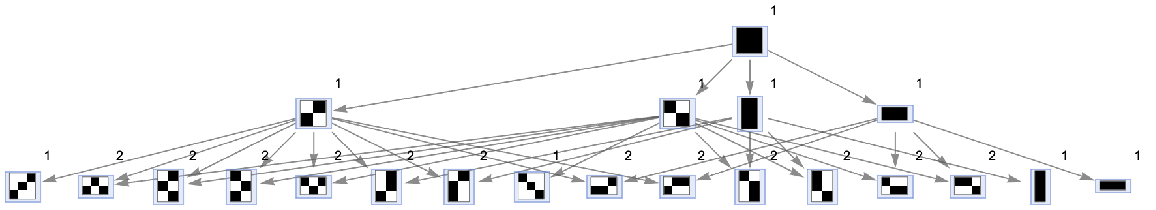 |
Show the structure of the graph, without labels:
| In[29]:= | ![ResourceFunction[
"MultiwayAggregationSystem"][<|"OuterTotalisticCode" -> 4, "Dimension" -> 2|>, CenterArray[1, {1, 1}], 2, "StatesGraphStructure", "IncludeStatePathWeights" -> True, VertexLabels -> "VertexWeight"]](https://www.wolframcloud.com/obj/resourcesystem/images/fa5/fa59000f-249d-45f0-bd3b-38f71609dbde/70997e04b2a4980d.png) |
| Out[29]= | 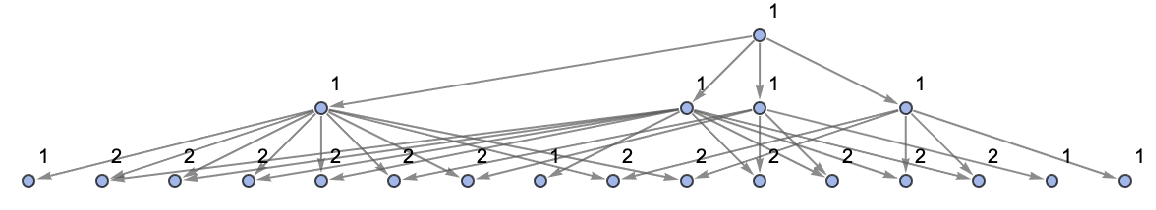 |
Run outer totalistic code 4 using the default (Moore) neighborhood:
| In[30]:= |
| Out[30]= | 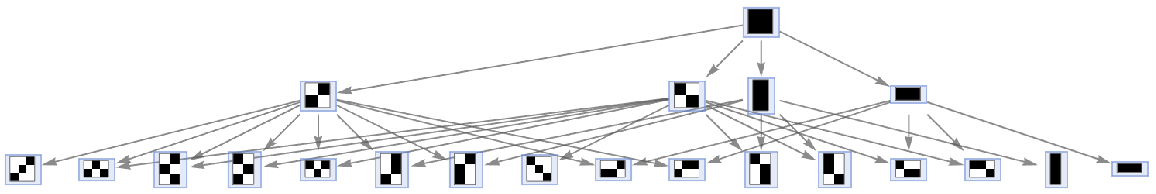 |
Run the same outer totalistic rule using the von Neumann neighborhood instead:
| In[31]:= |
| Out[31]= | 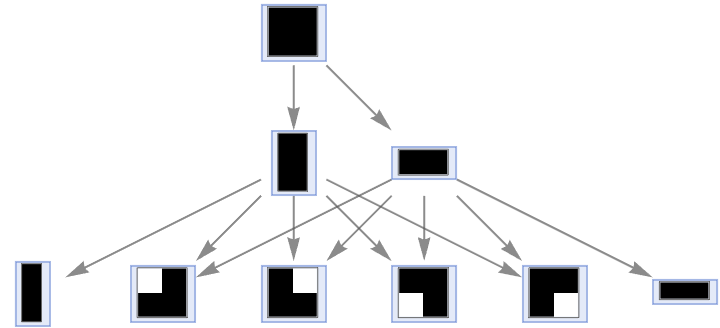 |
Simulate a multiway aggregation system with three colors:
| In[32]:= |
| Out[32]= | 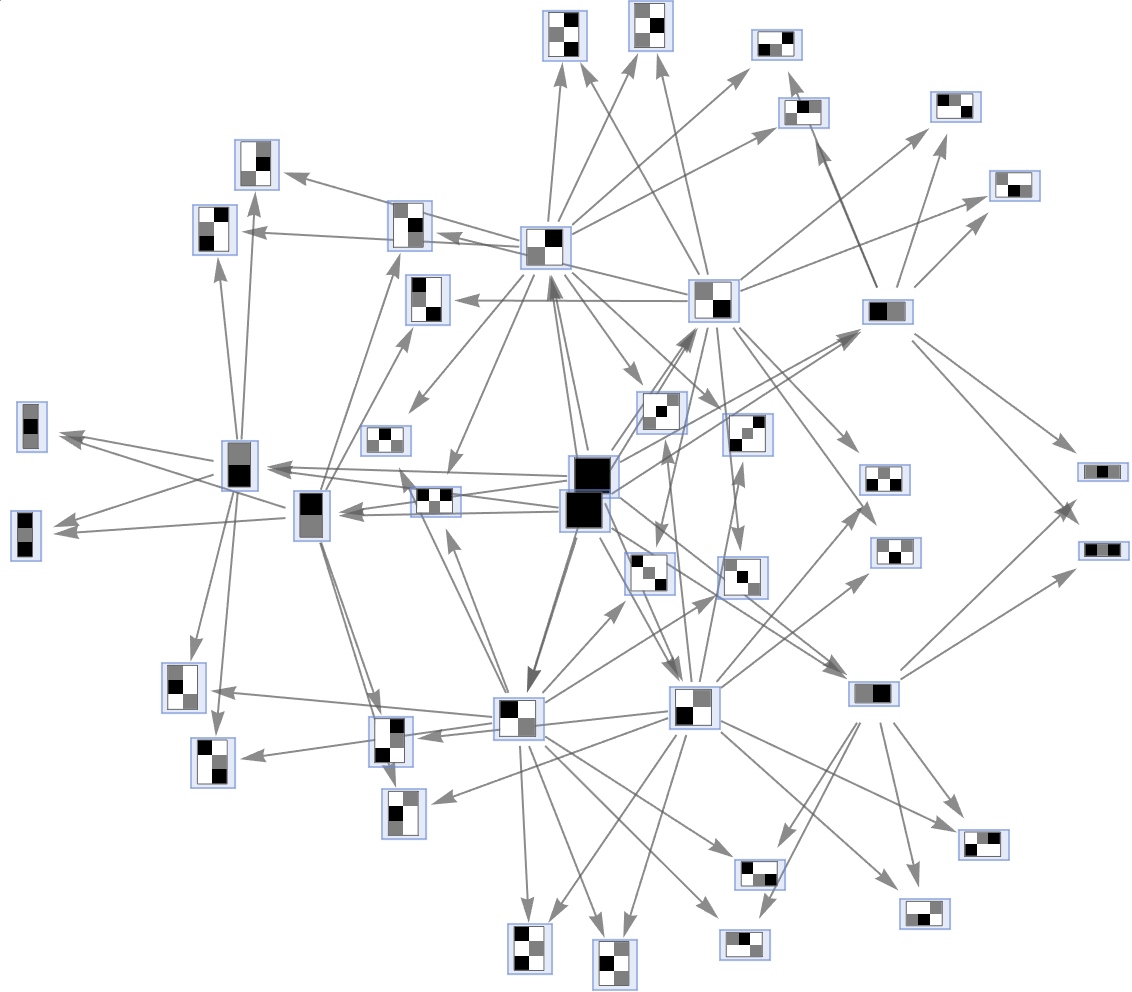 |
Simulate a multiway aggregation system specified using growth cases:
| In[33]:= |
| Out[33]= | 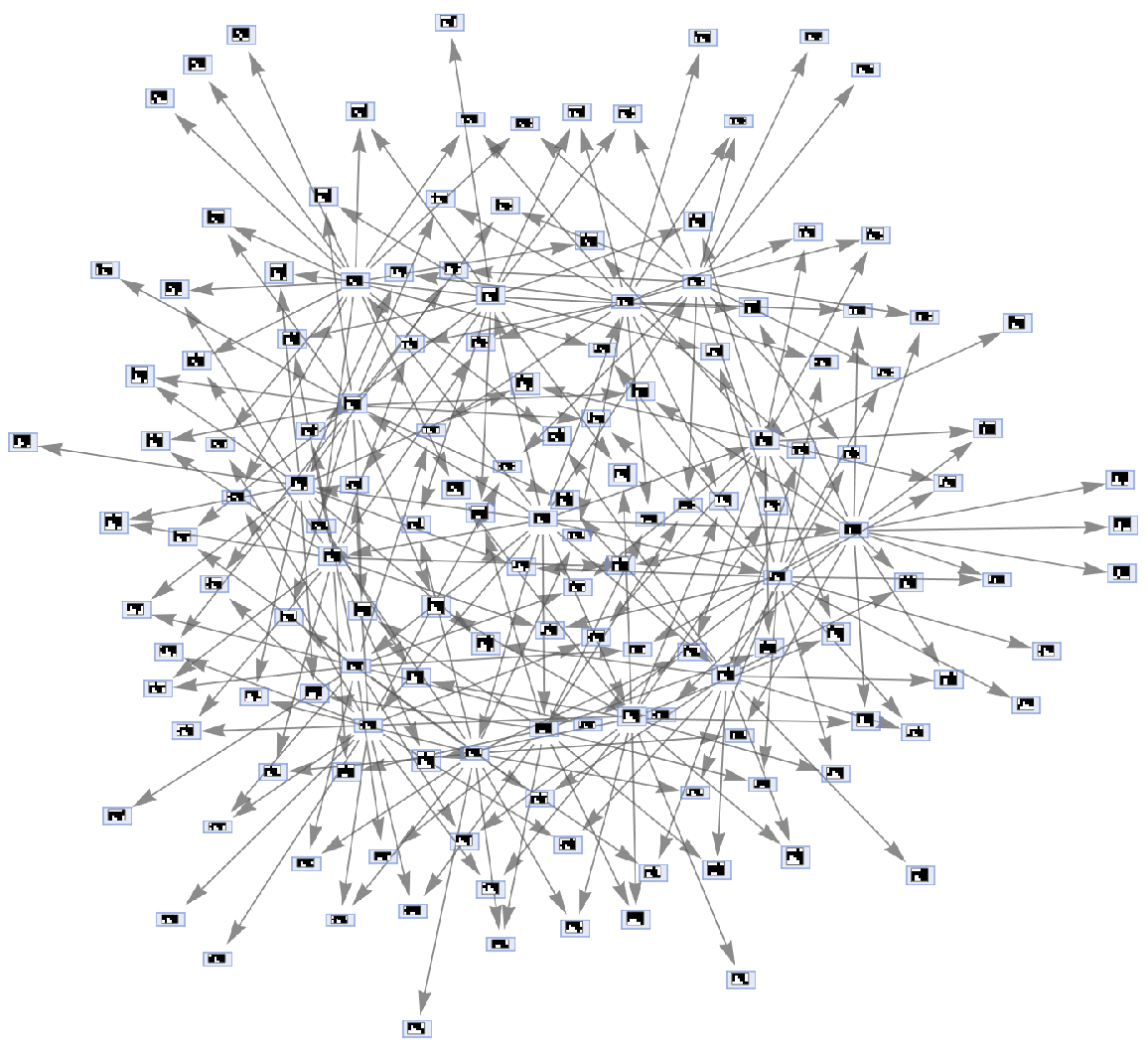 |
MultiwayAggregationSystem accepts both individual initial conditions and lists of initial conditions:
| In[34]:= |
| Out[34]= | 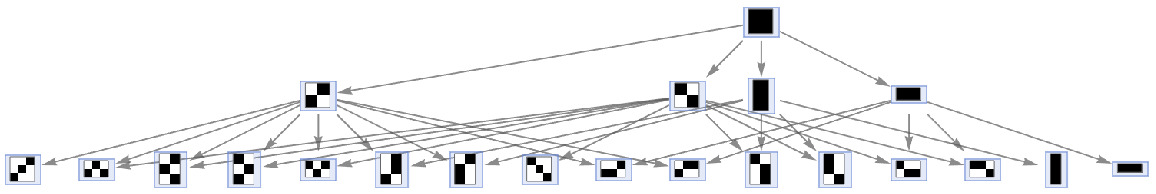 |
| In[35]:= |
| Out[35]= | 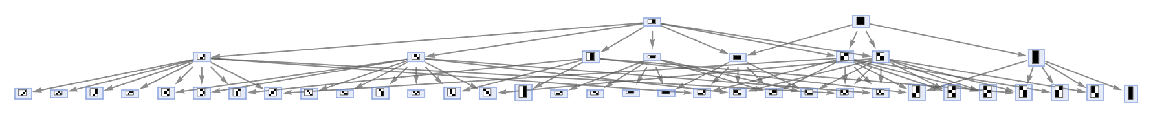 |
Apply only the first possible event at each step:
| In[36]:= |
| Out[36]= |
Apply the first and last possible events at each step:
| In[37]:= |
| Out[37]= |  |
By default, states are labeled by their contents:
| In[38]:= |
| Out[38]= | 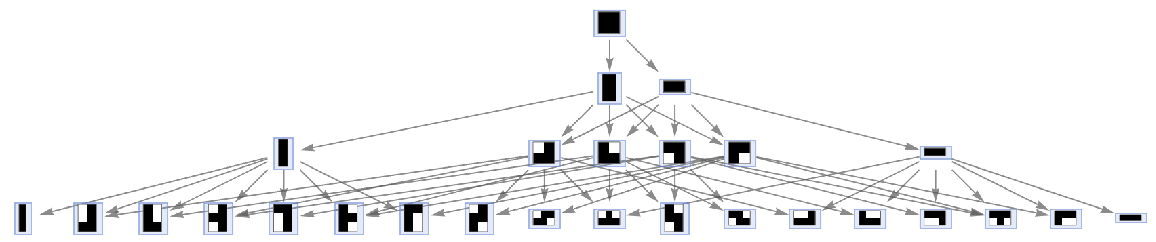 |
Use no labeling for states:
| In[39]:= | ![ResourceFunction[
"MultiwayAggregationSystem"][<|"OuterTotalisticCode" -> 4, "Dimension" -> 2, "Neighborhood" -> 5|>, CenterArray[1, {1, 1}], 3, "StatesGraph", "StateRenderingFunction" -> None]](https://www.wolframcloud.com/obj/resourcesystem/images/fa5/fa59000f-249d-45f0-bd3b-38f71609dbde/1f5d2ee0c282d1af.png) |
| Out[39]= | 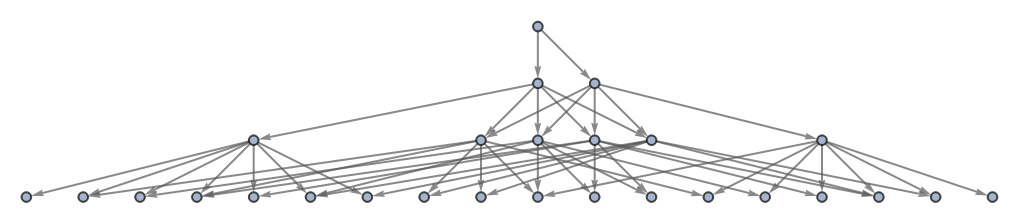 |
"StatesGraphStructure" yields the same result:
| In[40]:= |
| Out[40]= | 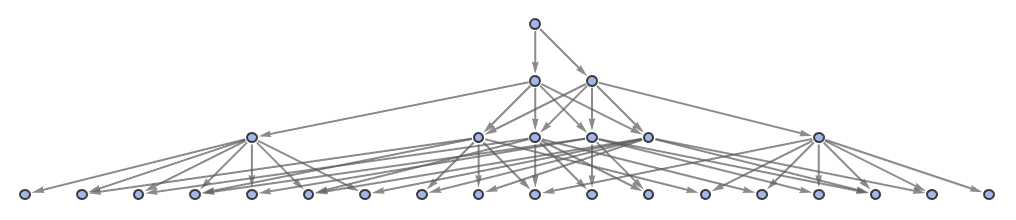 |
Use raw state names as node labels:
| In[41]:= | ![ResourceFunction[
"MultiwayAggregationSystem"][<|"OuterTotalisticCode" -> 4, "Dimension" -> 2, "Neighborhood" -> 5|>, CenterArray[1, {1, 1}], 3, "StatesGraph", "StateRenderingFunction" -> Inherited]](https://www.wolframcloud.com/obj/resourcesystem/images/fa5/fa59000f-249d-45f0-bd3b-38f71609dbde/44d73f2e1c25a25a.png) |
| Out[41]= | 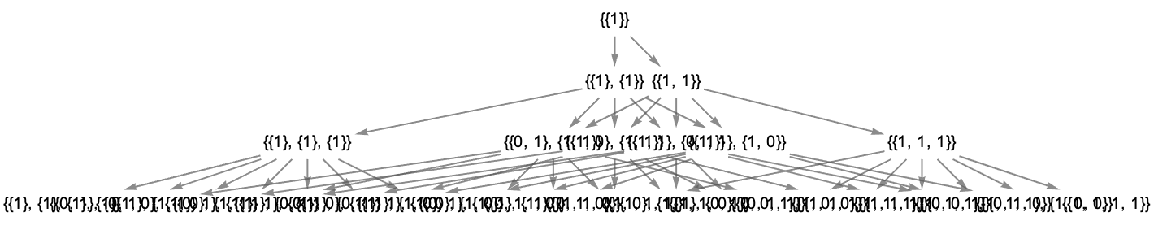 |
Use a named shape as each state label:
| In[42]:= | ![ResourceFunction[
"MultiwayAggregationSystem"][<|"OuterTotalisticCode" -> 4, "Dimension" -> 2, "Neighborhood" -> 5|>, CenterArray[1, {1, 1}], 3, "StatesGraph", "StateRenderingFunction" -> "Square"]](https://www.wolframcloud.com/obj/resourcesystem/images/fa5/fa59000f-249d-45f0-bd3b-38f71609dbde/165aabb72ef6a291.png) |
| Out[42]= | 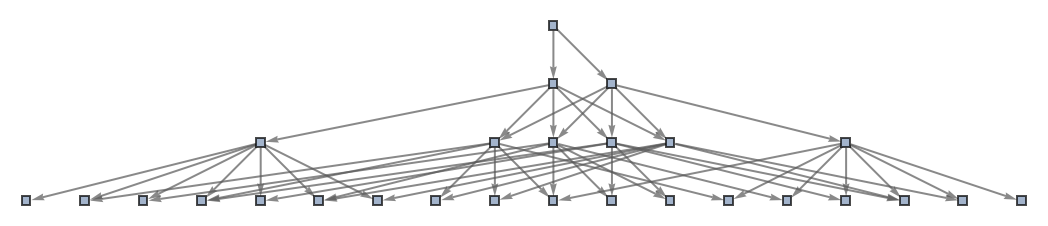 |
By default, both states and events are labeled by their contents:
| In[43]:= |
| Out[43]= | 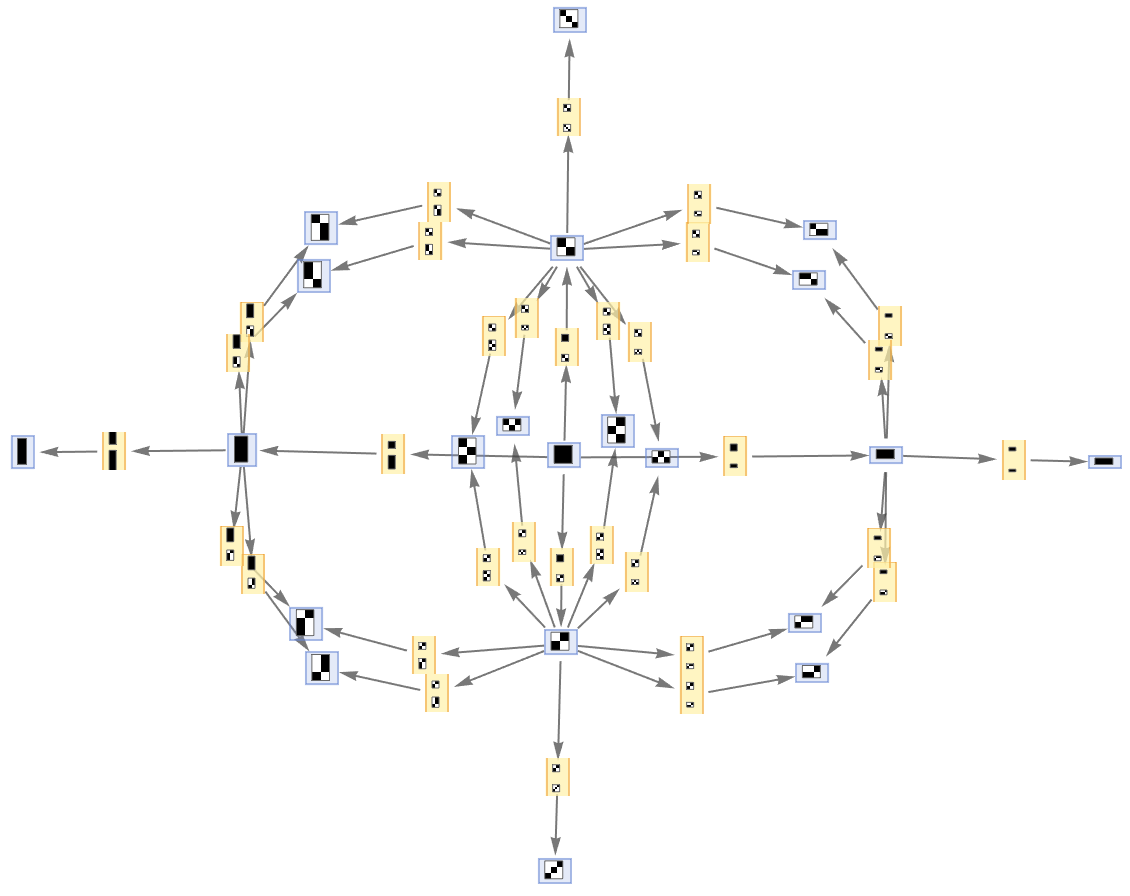 |
Use no labels for states or events:
| In[44]:= | ![ResourceFunction[
"MultiwayAggregationSystem"][<|"OuterTotalisticCode" -> 4, "Dimension" -> 2|>, CenterArray[1, {1, 1}], 2, "EvolutionEventsGraph", "StateRenderingFunction" -> None, "EventRenderingFunction" -> None]](https://www.wolframcloud.com/obj/resourcesystem/images/fa5/fa59000f-249d-45f0-bd3b-38f71609dbde/48a59652bdd739e8.png) |
| Out[44]= | 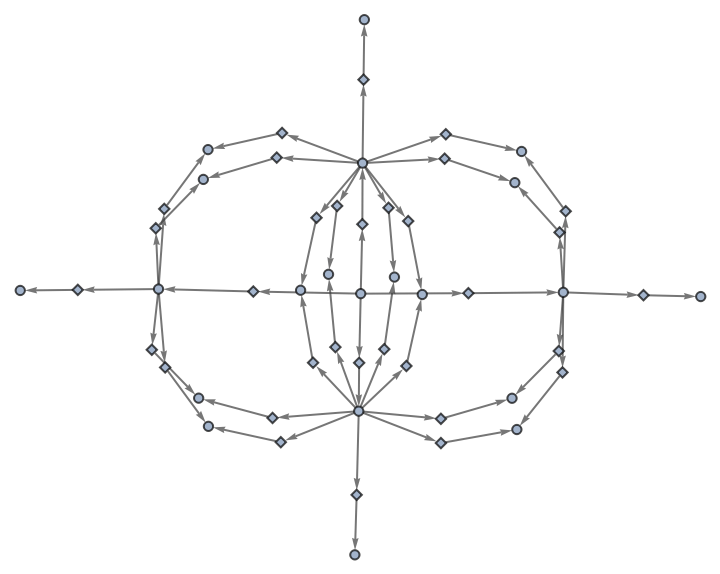 |
"EvolutionEventsGraphStructure" yields an equivalent result:
| In[45]:= |
| Out[45]= | 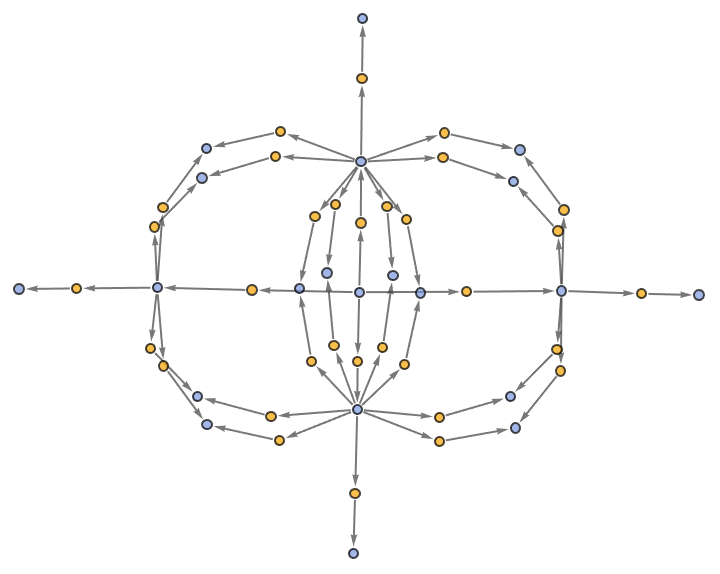 |
Use raw event expressions as their labels:
| In[46]:= | ![ResourceFunction[
"MultiwayAggregationSystem"][<|"OuterTotalisticCode" -> 4, "Dimension" -> 2|>, CenterArray[1, {1, 1}], 2, "EvolutionEventsGraph", "StateRenderingFunction" -> None, "EventRenderingFunction" -> Inherited]](https://www.wolframcloud.com/obj/resourcesystem/images/fa5/fa59000f-249d-45f0-bd3b-38f71609dbde/7a11b4f0ce7c01fb.png) |
| Out[46]= | 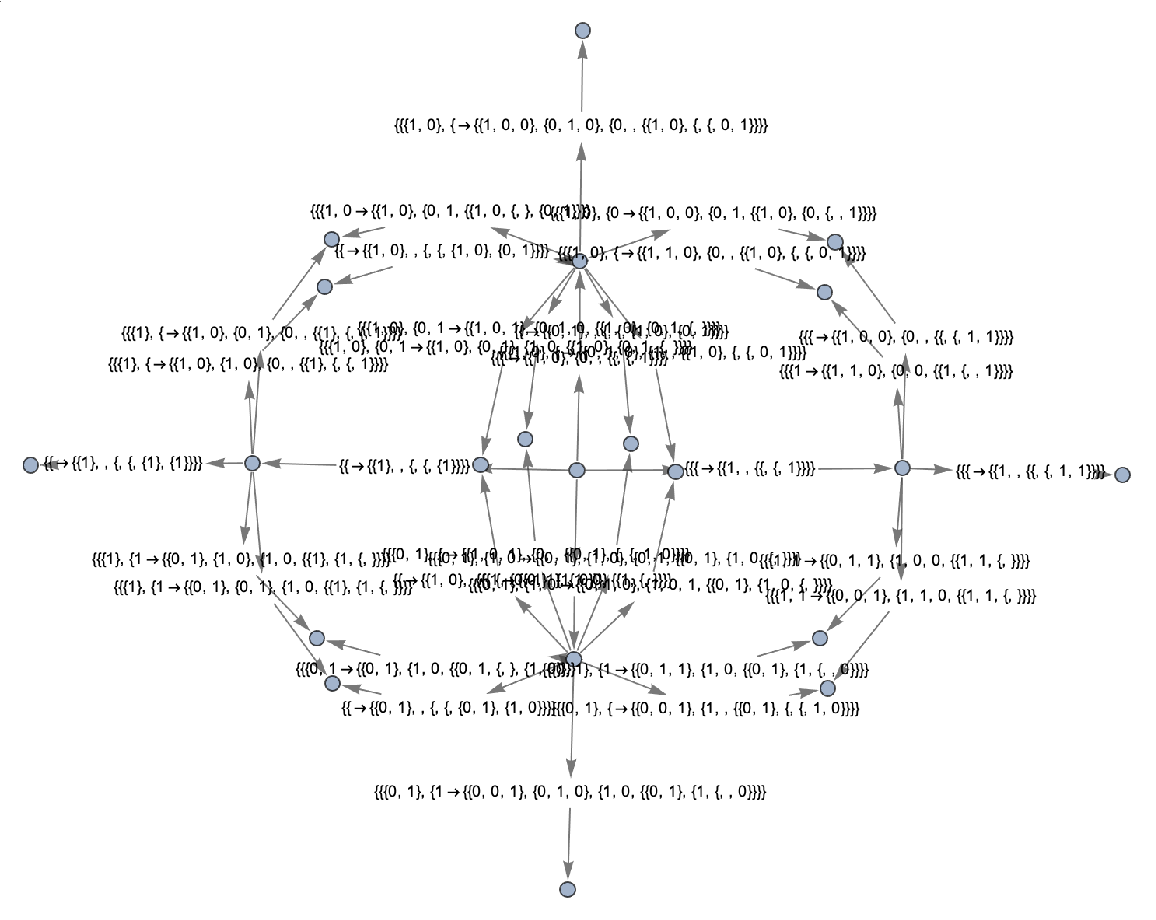 |
By default, "AllEventsList" does not include initialization events:
| In[47]:= |
| Out[47]= |  |
The option "IncludeInitializationEvents" allows one to override this default:
| In[48]:= |
| Out[48]= |  |
Initialization events have special default rendering:
| In[49]:= | ![ResourceFunction[
"MultiwayAggregationSystem"][<|"OuterTotalisticCode" -> 466, "Dimension" -> 2|>, {{{1, 0, 1}}}, 2, "EvolutionEventsGraph", "IncludeInitializationEvents" -> True, VertexSize -> 1]](https://www.wolframcloud.com/obj/resourcesystem/images/fa5/fa59000f-249d-45f0-bd3b-38f71609dbde/412fd3d84920881a.png) |
| Out[49]= | 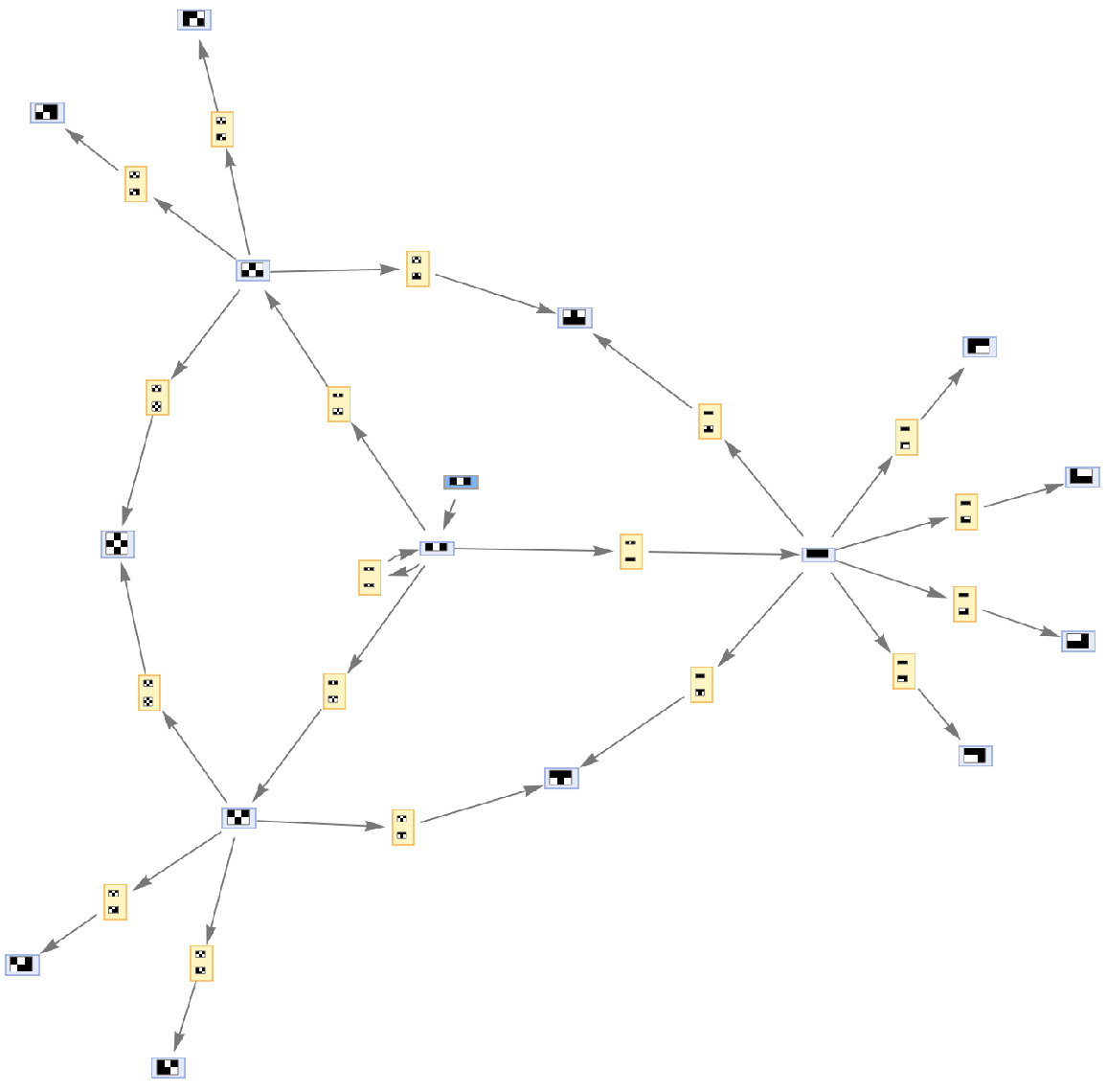 |
Place arrows in the middle of edges:
| In[50]:= | ![ResourceFunction[
"MultiwayAggregationSystem"][<|"OuterTotalisticCode" -> 4, "Dimension" -> 2, "Neighborhood" -> 5|>, 3, "StatesGraph", EdgeShapeFunction -> GraphElementData["ShortFilledArrow", "ArrowSize" -> 0.03], VertexSize -> 1]](https://www.wolframcloud.com/obj/resourcesystem/images/fa5/fa59000f-249d-45f0-bd3b-38f71609dbde/3fe5aa33174adfcd.png) |
| Out[50]= | 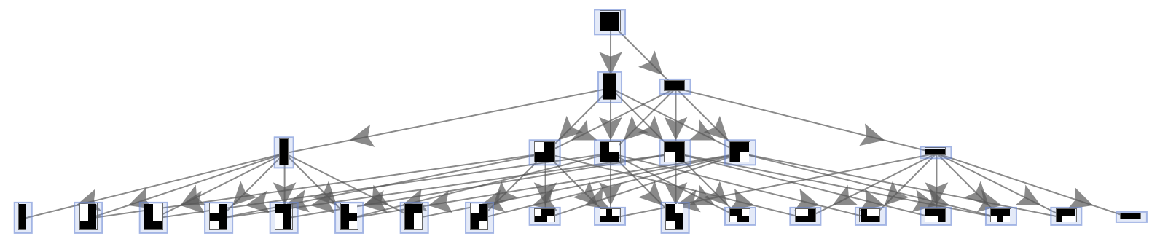 |
Generate an example of multiway aggregation system evolution:
| In[51]:= |
| Out[51]= | 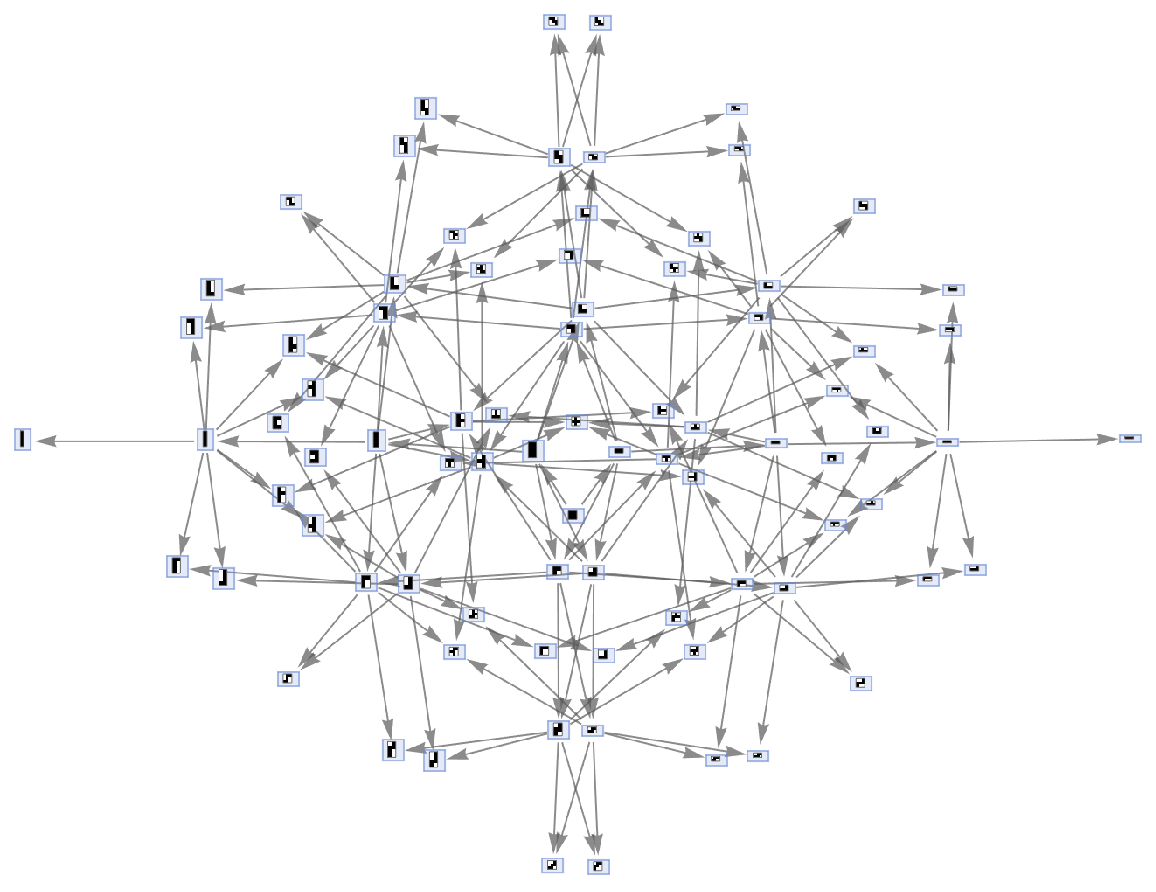 |
Force a layered digraph embedding:
| In[52]:= | ![ResourceFunction[
"MultiwayAggregationSystem"][<|"OuterTotalisticCode" -> 4, "Dimension" -> 2, "Neighborhood" -> 5|>, 4, "StatesGraph", GraphLayout -> "LayeredDigraphEmbedding", AspectRatio -> 1/2, VertexSize -> 2]](https://www.wolframcloud.com/obj/resourcesystem/images/fa5/fa59000f-249d-45f0-bd3b-38f71609dbde/5b071c5fda93bfaf.png) |
| Out[52]= | 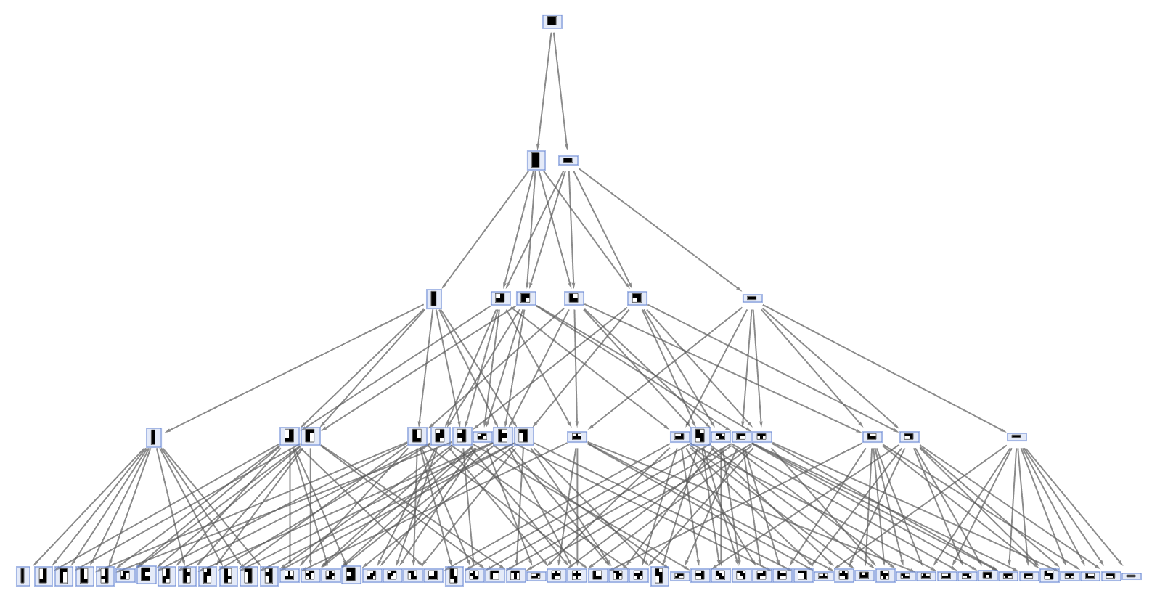 |
By default, equivalent states are merged across all time steps:
| In[53]:= |
| Out[53]= | 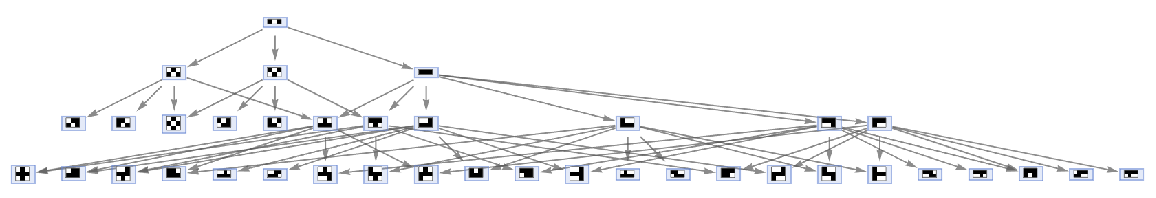 |
| In[54]:= |
| Out[54]= |  |
Merging of equivalent states across different time steps can be prevented by including step numbers:
| In[55]:= |
| Out[55]= | 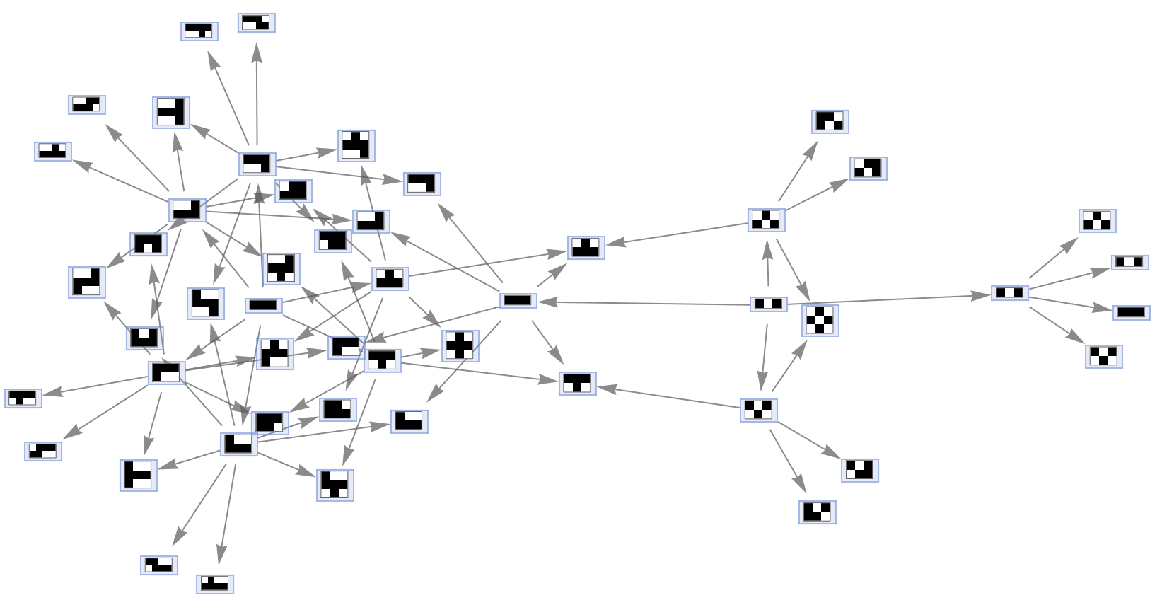 |
| In[56]:= |
| Out[56]= |  |
Merging of equivalent states at the same time step can be prevented by also including state IDs:
| In[57]:= | ![ResourceFunction[
"MultiwayAggregationSystem"][<|"OuterTotalisticCode" -> 466, "Dimension" -> 2|>, {{{1, 0, 1}}}, 2, "StatesGraph", "IncludeStepNumber" -> True, "IncludeStateID" -> True, VertexSize -> 1]](https://www.wolframcloud.com/obj/resourcesystem/images/fa5/fa59000f-249d-45f0-bd3b-38f71609dbde/15812ac40b7d6250.png) |
| Out[57]= | 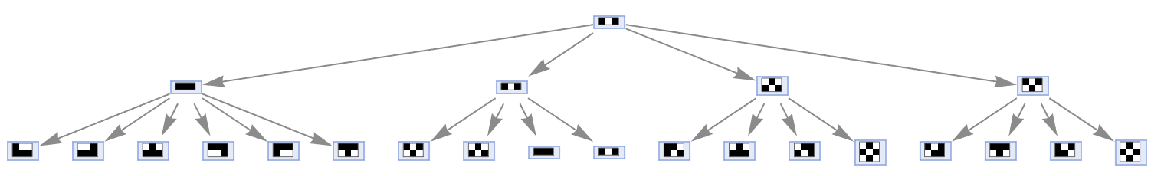 |
| In[58]:= |
| Out[58]= |  |
Step numbers and IDs also apply to events:
| In[59]:= | ![ResourceFunction[
"MultiwayAggregationSystem"][<|"OuterTotalisticCode" -> 466, "Dimension" -> 2|>, {{{1, 0, 1}}}, 2, "EvolutionEventsGraph", "IncludeStepNumber" -> True, "IncludeStateID" -> True, VertexSize -> 1]](https://www.wolframcloud.com/obj/resourcesystem/images/fa5/fa59000f-249d-45f0-bd3b-38f71609dbde/467d213cbf75a0df.png) |
| Out[59]= | 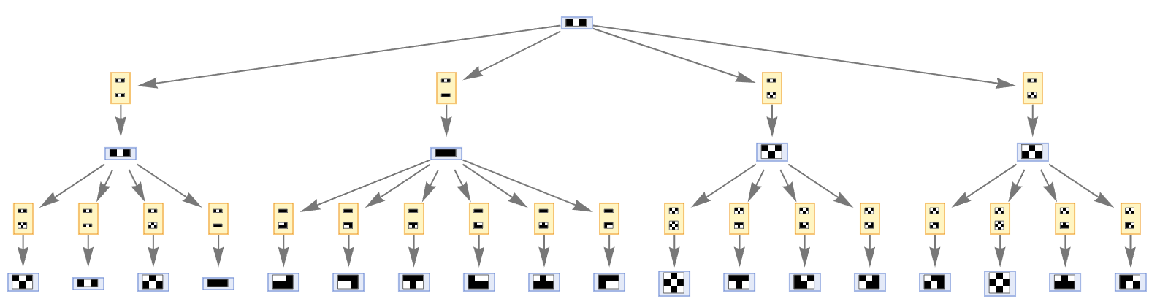 |
See the events:
| In[60]:= |
| Out[60]= |  |
By default, multiple instances of equivalent updating events are merged in the states graph:
| In[61]:= |
| Out[61]= | 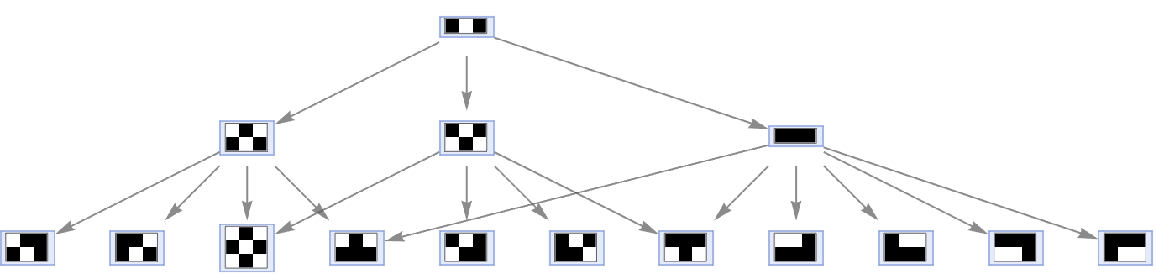 |
Merging of equivalent events can be prevented by including event instances:
| In[62]:= |
| Out[62]= | 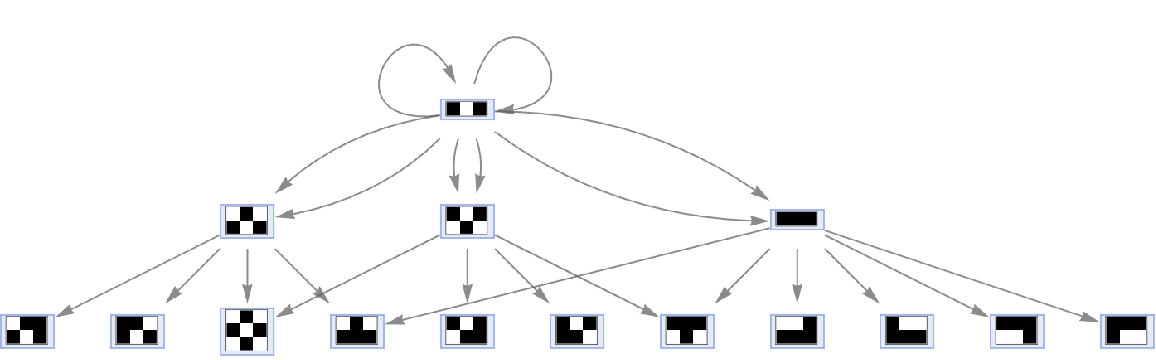 |
Vertices of a states graph can be weighted by their relative rate of occurrence at each time step:
| In[63]:= | ![ResourceFunction[
"MultiwayAggregationSystem"][<|"OuterTotalisticCode" -> 4, "Dimension" -> 2|>, CenterArray[1, {1, 1}], 2, "StatesGraph", "IncludeStateWeights" -> True, VertexLabels -> "VertexWeight", VertexSize -> 1]](https://www.wolframcloud.com/obj/resourcesystem/images/fa5/fa59000f-249d-45f0-bd3b-38f71609dbde/7ba6ae1621e04fee.png) |
| Out[63]= | 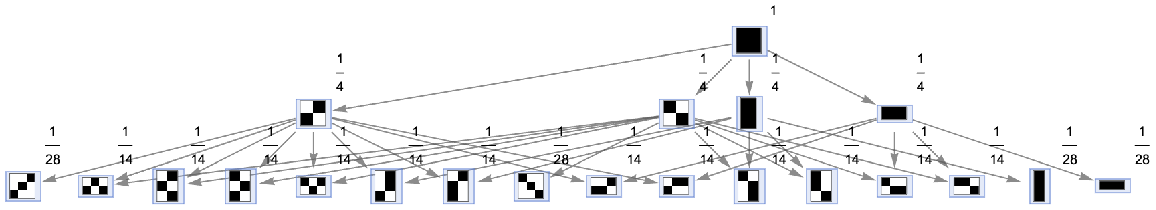 |
Vertices can also be weighted by the number of distinct evolution paths that lead to them:
| In[64]:= | ![ResourceFunction[
"MultiwayAggregationSystem"][<|"OuterTotalisticCode" -> 4, "Dimension" -> 2|>, CenterArray[1, {1, 1}], 2, "StatesGraph", "IncludeStatePathWeights" -> True, VertexLabels -> "VertexWeight", VertexSize -> 1]](https://www.wolframcloud.com/obj/resourcesystem/images/fa5/fa59000f-249d-45f0-bd3b-38f71609dbde/3d601f700f65b4d8.png) |
| Out[64]= | 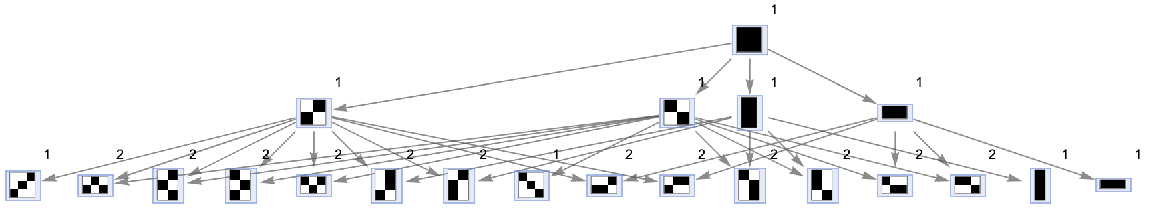 |
By default, "CausalGraphInstances" returns all possible causal graphs:
| In[65]:= | ![Graph[#, VertexSize -> 1] & /@ Take[ResourceFunction[
"MultiwayAggregationSystem"][<|"OuterTotalisticCode" -> 4, "Dimension" -> 2, "Neighborhood" -> 5|>, CenterArray[1, {1, 1}], 3, "CausalGraphInstances"], 10]](https://www.wolframcloud.com/obj/resourcesystem/images/fa5/fa59000f-249d-45f0-bd3b-38f71609dbde/36cbe03c00e8b689.png) |
| Out[65]= | 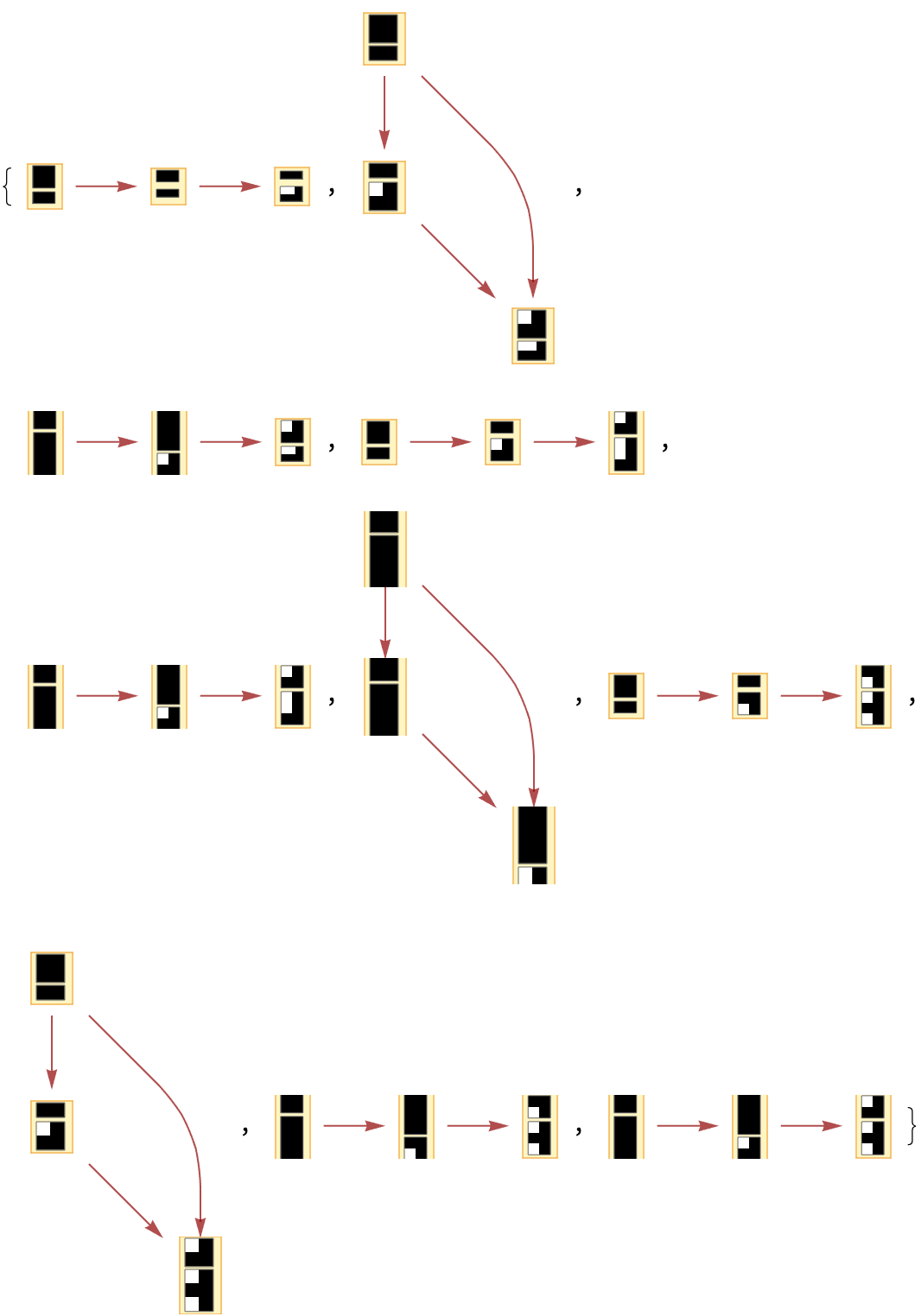 |
The number of causal graphs returned can be limited using MaxItems:
| In[66]:= | ![Graph[#, VertexSize -> 1] & /@ ResourceFunction[
"MultiwayAggregationSystem"][<|"OuterTotalisticCode" -> 4, "Dimension" -> 2, "Neighborhood" -> 5|>, CenterArray[1, {1, 1}], 3,
"CausalGraphInstances", MaxItems -> 5]](https://www.wolframcloud.com/obj/resourcesystem/images/fa5/fa59000f-249d-45f0-bd3b-38f71609dbde/4af2aa470927c896.png) |
| Out[66]= | 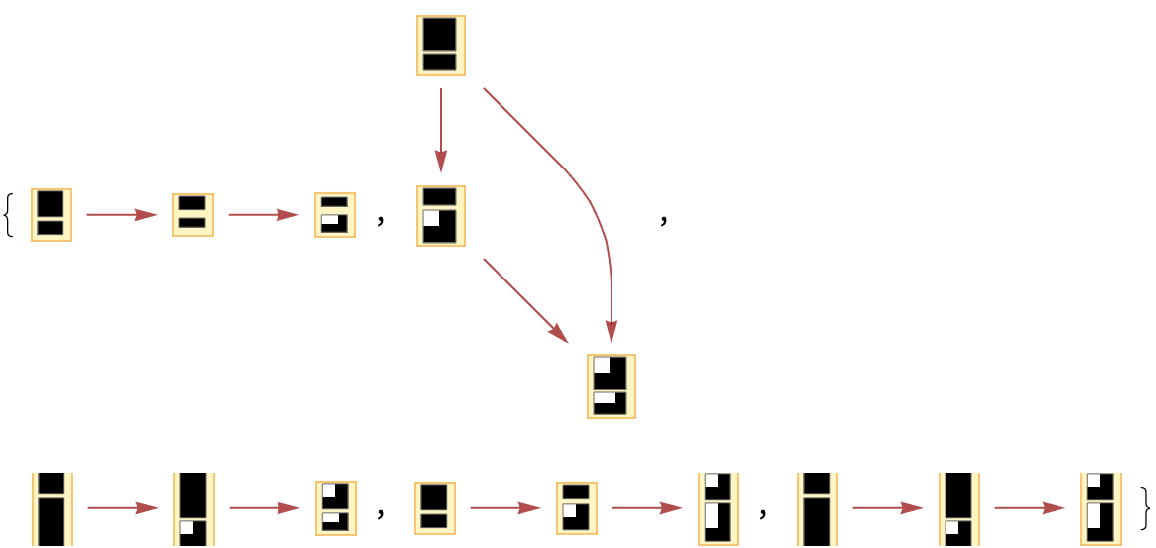 |
By default, "BranchPairsList" returns only a list of branch pairs:
| In[67]:= |
| Out[67]= |  |
Common predecessor states can be shown using "GivePredecessors":
| In[68]:= |
| Out[68]= |  |
Similarly, "BranchPairResolutionsList" by default lists only resolved and unresolved branch pairs:
| In[69]:= |
| Out[69]= |  |
Common resolvents of resolved branch pairs can be shown using "GiveResolvents":
| In[70]:= |
| Out[70]= |  |
Show both common predecessors and common resolvents, where appropriate:
| In[71]:= | ![ResourceFunction[
"MultiwayAggregationSystem"][<|"OuterTotalisticCode" -> 466, "Dimension" -> 2|>, {{{1, 0, 1}}}, 2, "BranchPairResolutionsList", "GivePredecessors" -> True, "GiveResolvents" -> True]](https://www.wolframcloud.com/obj/resourcesystem/images/fa5/fa59000f-249d-45f0-bd3b-38f71609dbde/5287152c516281d1.png) |
| Out[71]= |  |
This work is licensed under a Creative Commons Attribution 4.0 International License