Wolfram Language Paclet Repository
Community-contributed installable additions to the Wolfram Language
An integrated Mathematica framework for computing residue-level contacts in oligomeric protein assemblies
Contributed by: Yingying Jin
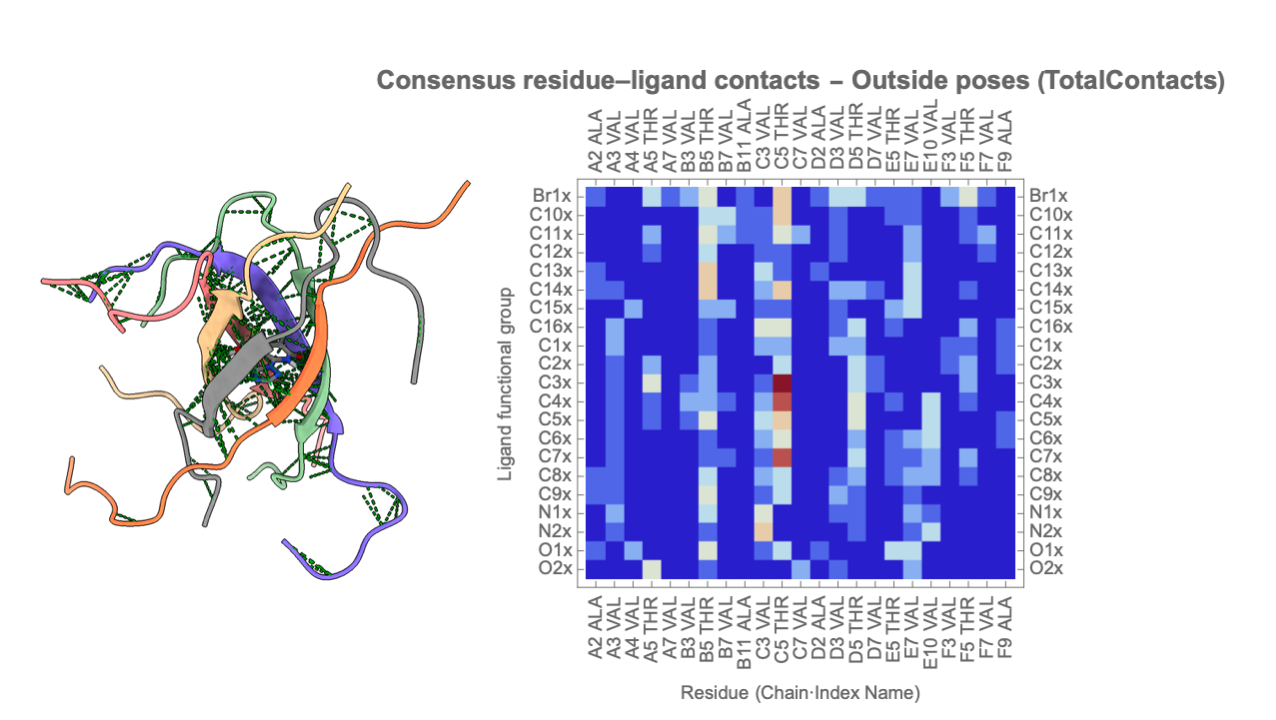
ProteinLigandInteractionAnalysis is a Mathematica paclet designed to provide a complete, reproducible framework for quantitative analysis of protein–ligand interactions, particularly within oligomeric and β-barrel–like assemblies such as NACore hexamers. The paclet offers an end-to-end workflow that includes automated PDB parsing, extraction of barrel geometry parameters, ligand functional-group annotation, computation of residue-level contact statistics, and categorical pose classification (Inside / Borderline / Outside). Its modular architecture standardizes core metrics—such as radial and axial ligand COM positions, pore-occupancy fractions, and contact-based interaction fingerprints—enabling rapid comparison across large pose ensembles like 50-structure docking outputs. In addition, ProteinLigandInteractionAnalysis provides powerful visualization functions for annotated molecular renderings, consensus heatmaps, and curated pose summaries, making it easy to evaluate binding trends across multiple ligands or sequence variants. Together, these tools form a flexible and extensible platform for high-resolution structure-based interpretation of protein–ligand binding behavior.
To install this paclet in your Wolfram Language environment,
evaluate this code:
PacletInstall["YingyingJin/ProteinLigandInteractionAnalysis"]
To load the code after installation, evaluate this code:
Needs["YingyingJin`ProteinLigandInteractionAnalysis`"]
Define Protein Structure:
| In[1]:= | ![cylFile = "/Users/yingyingjin/Dropbox/NewMoleculeProject/DynamicBind/cylindrinA.pdb";
barrelParams = ComputeProteinParameters[cylFile];
barrelParams](https://www.wolframcloud.com/obj/resourcesystem/images/d36/d366f361-93d8-4b3a-9edd-b74ed5d350cc/1865823444e5a252.png) |
| Out[3]= |
| In[4]:= | ![(*Assume barrelParams already exists*)
DrawBarrelGeometry[barrelParams_Association, finInside_ : 0.30,(*used in LabelPose,shown in label text*)
finBorder_ : 0.15,(*used in LabelPose,shown in label text*)
rSlack_ : 0.5 (*radial slack used for borderline region*)] := Module[{rPore, zLow, zHigh, zMid, rMax, dz, rBorder}, {rPore, zLow, zHigh} = Lookup[barrelParams, {"Rpore", "Zlow", "Zhigh"}];
zMid = (zLow + zHigh)/2.;
dz = zHigh - zLow;
rBorder = rPore + rSlack;
rMax = 1.2 rBorder;(*extend a bit past the borderline region*)
Graphics[{(*1) Overall barrel "shell" region*){Lighter[Gray, 0.9], EdgeForm[Red], Rectangle[{-rMax, zLow}, {rMax, zHigh}]},(*2) Strongly INSIDE region:
r\[LessEqual]Rpore*){Directive[Lighter[Blue, 0.85], Opacity[0.5]],
Rectangle[{-rPore, zLow}, {rPore, zHigh}]},(*3) Borderline radial band:Rpore<r\[LessEqual]Rpore+
rSlack*){Directive[Lighter[Blue, 0.95], Opacity[0.5]], Rectangle[{-rBorder, zLow}, {-rPore, zHigh}], Rectangle[{rPore, zLow}, {rBorder, zHigh}]},(*4) Vertical lines marking Rpore and Rpore+
rSlack*){Blue, Thick, Dashed, Line[{{rPore, zLow}, {rPore, zHigh}}],
Line[{{-rPore, zLow}, {-rPore, zHigh}}], Line[{{rBorder, zLow}, {rBorder, zHigh}}], Line[{{-rBorder, zLow}, {-rBorder, zHigh}}]}, Text[Style["Rpore", 12, Blue], {rPore, zHigh + 0.06 dz}], Text[Style["Rpore + rSlack", 11, Blue], {rBorder, zHigh + 0.06 dz}],(*5) Pore radius at mid-
plane (same as before)*){Blue, Thick, Line[{{0, zMid}, {rPore, zMid}}], PointSize[0.015], Point[{0, zMid}], Text[Style["Rpore", 14, Blue], {rPore/2, zMid + 0.08 dz}]},(*6) Zlow and Zhigh planes*){Red, Thick, Line[{{-rMax, zLow}, {rMax, zLow}}], Line[{{-rMax, zHigh}, {rMax, zHigh}}], Text[Style["Zlow", 13, Red], {rMax*1.05, zLow}], Text[Style["Zhigh", 13, Red], {rMax*1.05, zHigh}]},(*7) z-
axis*){Black, Arrow[{{0, zLow - 0.25 dz}, {0, zHigh + 0.25 dz}}], Text[Style["z-axis", 12], {0, zHigh + 0.32 dz}]},(*8) Region annotations tied to LabelPose thresholds*)
Text[Style[
"Strongly inside region\n(r \[LessEqual] Rpore,\nfin \[GreaterEqual] " <> ToString[NumberForm[finInside, {2, 2}]] <> ")", 11, Blue], {0, zMid - 0.25 dz}], Text[Style[
"Borderline band\n(Rpore < r \[LessEqual] Rpore + rSlack,\nfin \[GreaterEqual] " <> ToString[NumberForm[finBorder, {2, 2}]] <> ")", 10, Darker[Blue]], {rBorder - rSlack/2., zMid + 0.25 dz}]}, Frame -> True, FrameLabel -> {"radial distance r", "position along axis z"}, AspectRatio -> 1.2, ImageSize -> 400]]](https://www.wolframcloud.com/obj/resourcesystem/images/d36/d366f361-93d8-4b3a-9edd-b74ed5d350cc/0a8573f46e3fd707.png) |
| In[5]:= |
| Out[5]= | 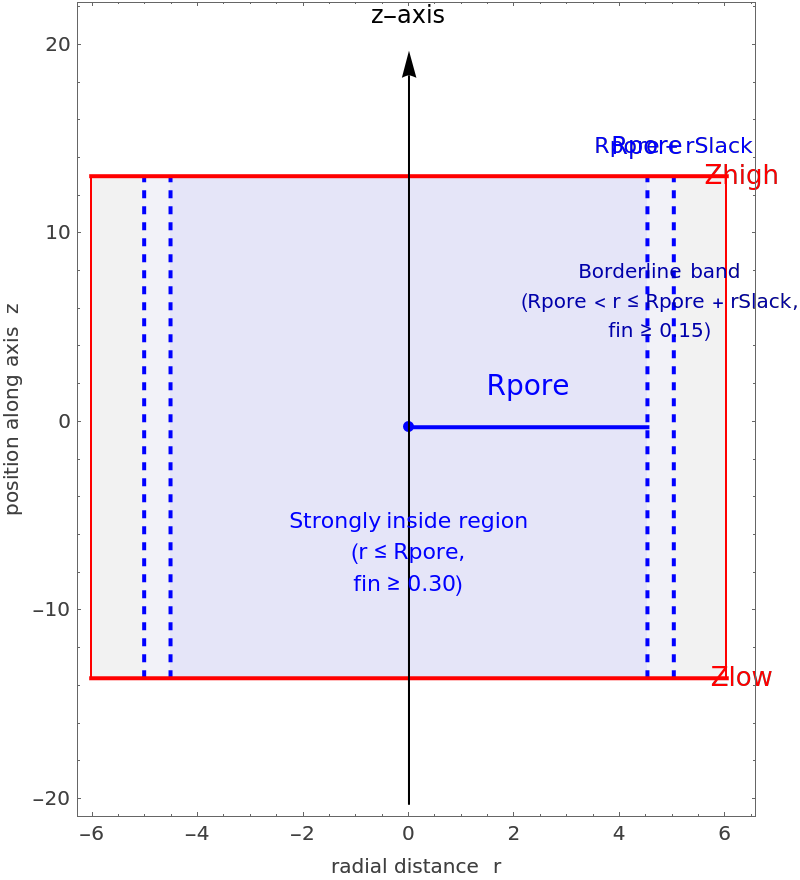 |
| In[6]:= |
| In[7]:= | ![classified = Association@
Table[FileBaseName[f] -> ClassifyProteinLigandPose[f, barrelParams], {f, files}];](https://www.wolframcloud.com/obj/resourcesystem/images/d36/d366f361-93d8-4b3a-9edd-b74ed5d350cc/53ba24f945482df4.png) |
| In[8]:= | ![(*Get keys corresponding to inside poses*)
insideKeys = Keys@Select[classified, #Label == "Inside" &];
(*Extract numeric rank part*)
insideRanksZINC1 = ToExpression@StringReplace[insideKeys, "rank_" -> ""]
(*Build file paths*)
insideFiles = FileNameJoin[{pdbDir, "rank_" <> ToString[#] <> ".pdb"}] & /@ insideRanksZINC1;
(*Import PDBs (or do further analysis)*)
insidePDBsZINC1 = Import /@ insideFiles](https://www.wolframcloud.com/obj/resourcesystem/images/d36/d366f361-93d8-4b3a-9edd-b74ed5d350cc/380b72455a530a01.png) |
| Out[5]= |
| Out[9]= | 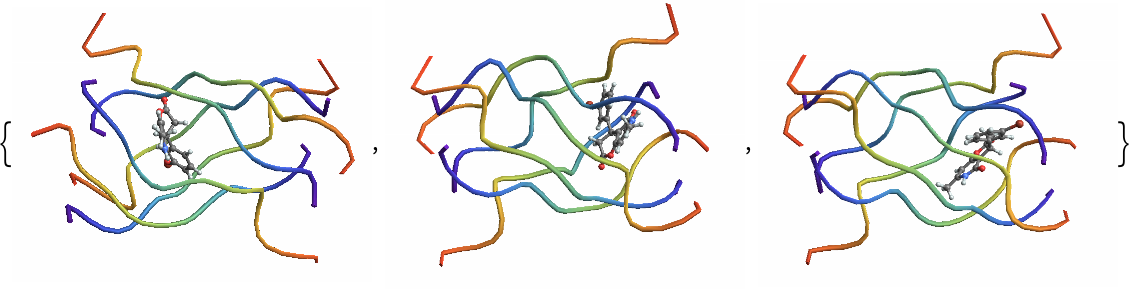 |
| In[10]:= | ![(*Get keys corresponding to inside poses*)
borderlineKeys = Keys@Select[classified, #Label == "Borderline" &];
(*Extract numeric rank part*)
borderlineRanksZINC1 = ToExpression@StringReplace[borderlineKeys, "rank_" -> ""]
(*Build file paths*)
borderlineFiles = FileNameJoin[{pdbDir, "rank_" <> ToString[#] <> ".pdb"}] & /@ borderlineRanksZINC1;
(*Import PDBs (or do further analysis)*)
borderlinePDBsZINC1 = Import /@ borderlineFiles](https://www.wolframcloud.com/obj/resourcesystem/images/d36/d366f361-93d8-4b3a-9edd-b74ed5d350cc/3d3be6d6a8f81e56.png) |
| Out[11]= |
| Out[13]= |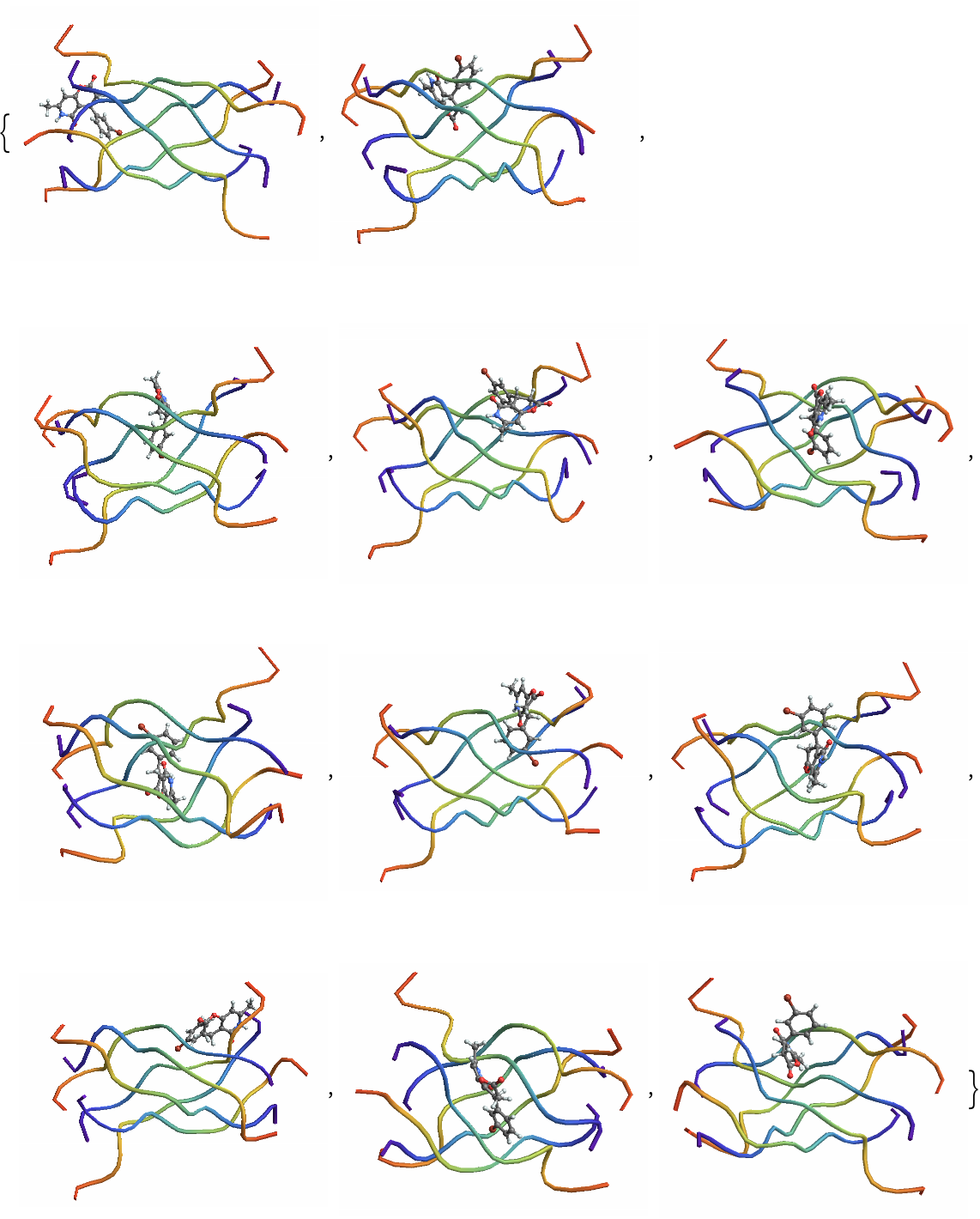  |
| In[14]:= | ![(*Get keys corresponding to inside poses*)
outsidelineKeys = Keys@Select[classified, #Label == "Outside" &];
(*Extract numeric rank part*)
outsidelineRanksZINC1 = ToExpression@StringReplace[outsidelineKeys, "rank_" -> ""]
(*Build file paths*)
outsidelineFiles = FileNameJoin[{pdbDir, "rank_" <> ToString[#] <> ".pdb"}] & /@ outsidelineRanksZINC1;
(*Import PDBs (or do further analysis)*)
outsidelinePDBsZINC1 = Import /@ outsidelineFiles](https://www.wolframcloud.com/obj/resourcesystem/images/d36/d366f361-93d8-4b3a-9edd-b74ed5d350cc/4ed9f9e775eac7ec.png) |
| Out[15]= |
| Out[17]= |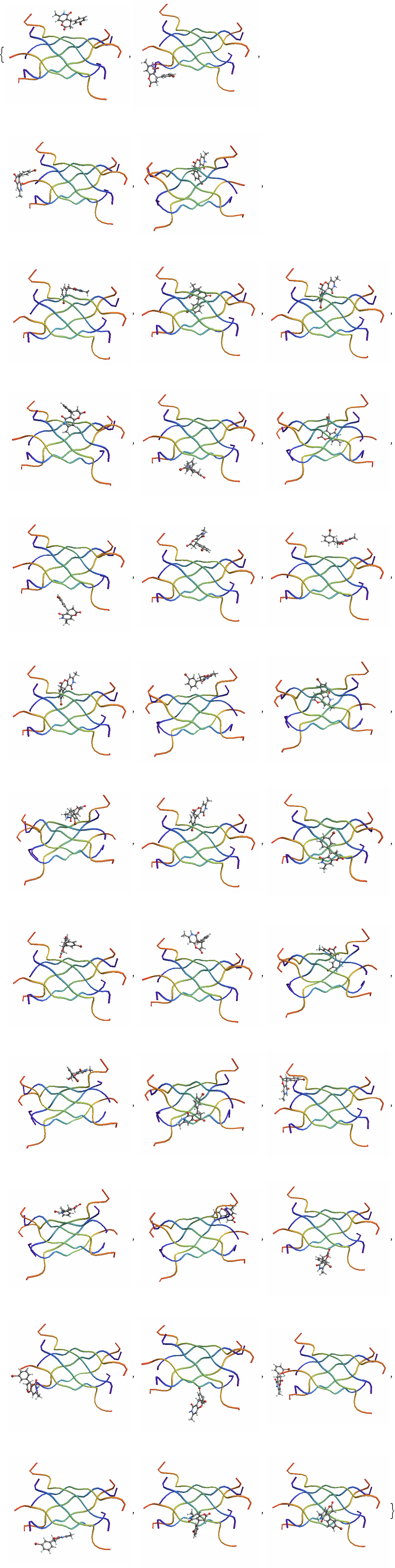  |
Wolfram Language Version 14.